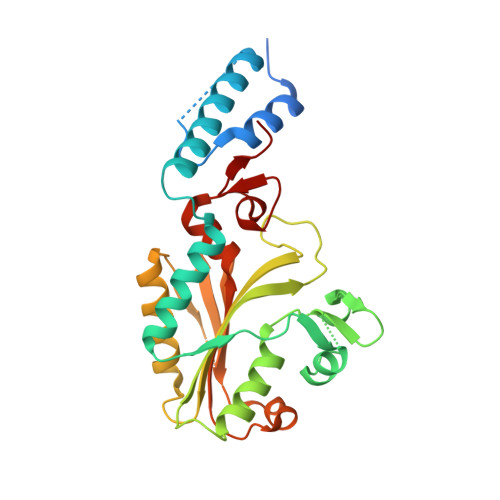
Structure of pyrrolysyl-tRNA synthetase, an archaeal enzyme for genetic code innovation.
Kavran, J.M., Gundllapalli, S., O'donoghue, P., Englert, M., Soll, D., Steitz, T.A.(2007) Proc Natl Acad Sci U S A 104: 11268-11273
- PubMed: 17592110
- DOI: https://doi.org/10.1073/pnas.0704769104
- Primary Citation of Related Structures:
2Q7E, 2Q7G, 2Q7H, 2ZIM - PubMed Abstract:
Pyrrolysine (Pyl), the 22nd natural amino acid and genetically encoded by UAG, becomes attached to its cognate tRNA by pyrrolysyl-tRNA synthetase (PylRS). We have determined three crystal structures of the Methanosarcina mazei PylRS complexed with either AMP-PNP, Pyl-AMP plus pyrophosphate, or the Pyl analogue N-epsilon-[(cylopentyloxy)carbonyl]-L-lysine plus ATP. The structures reveal that PylRS utilizes a deep hydrophobic pocket for recognition of the Pyl side chain. A comparison of these structures with previously determined class II tRNA synthetase complexes illustrates that different substrate specificities derive from changes in a small number of residues that form the substrate side-chain-binding pocket. The knowledge of these structures allowed the placement of PylRS in the aminoacyl-tRNA synthetase (aaRS) tree as the last known synthetase that evolved for genetic code expansion, as well as the finding that Pyl arose before the last universal common ancestral state. The PylRS structure provides an excellent framework for designing new aaRSs with altered amino acid specificity.
Organizational Affiliation:
Department of Molecular Biophysics and Biochemistry, Howard Hughes Medical Institute, Yale University, New Haven, CT 06520-8114, USA.