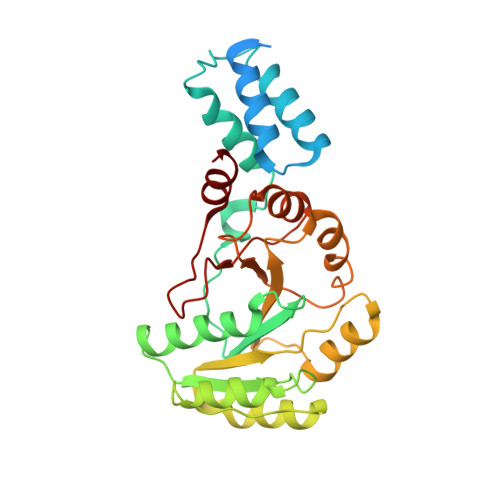
The crystal structure of the third signal-recognition particle GTPase FlhF reveals a homodimer with bound GTP.
Bange, G., Petzold, G., Wild, K., Parlitz, R.O., Sinning, I.(2007) Proc Natl Acad Sci U S A 104: 13621-13625
- PubMed: 17699634
- DOI: https://doi.org/10.1073/pnas.0702570104
- Primary Citation of Related Structures:
2PX0, 2PX3 - PubMed Abstract:
Flagella are well characterized as the organelles of locomotion and allow bacteria to react to environmental changes. The assembly of flagella is a multistep process and relies on a complex type III export machinery located in the cytoplasmic membrane. The FlhF protein is essential for the placement and assembly of polar flagella and has been classified as a signal-recognition particle (SRP)-type GTPase. SRP GTPases appeared early in evolution and form a unique subfamily within the guanine nucleotide binding proteins with only three members: the signal sequence-binding protein SRP54, the SRP receptor FtsY, and FlhF. We report the crystal structures of FlhF from Bacillus subtilis in complex with GTP and GMPPNP. FlhF shares SRP GTPase-specific features such as the presence of an N-terminal alpha-helical domain and the I-box insertion. It forms a symmetric homodimer sequestering a composite active site that contains two head-to-tail arranged nucleotides similar to the heterodimeric SRP-targeting complex. However, significant differences to the GTPases of SRP and the SRP receptor include the formation of a stable homodimer with GTP as well as severe modifications and even the absence of motifs involved in regulation of the other two SRP GTPases. Our results provide insights into SRP GTPases and their roles in two fundamentally different protein-targeting routes that both rely on efficient protein delivery to a secretion channel.
Organizational Affiliation:
Heidelberg University Biochemistry Center (BZH), INF 328, 69120 Heidelberg, Germany.