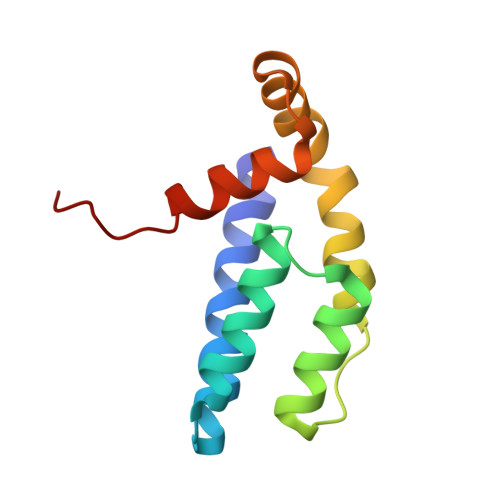
A TAF4-homology domain from the corepressor ETO is a docking platform for positive and negative regulators of transcription
Wei, Y., Liu, S., Lausen, J., Woodrell, C., Cho, S., Biris, N., Kobayashi, N., Wei, Y., Yokoyama, S., Werner, M.H.(2007) Nat Struct Mol Biol 14: 653-661
- PubMed: 17572682
- DOI: https://doi.org/10.1038/nsmb1258
- Primary Citation of Related Structures:
2PP4 - PubMed Abstract:
The eight twenty-one protein, ETO, is implicated in 12%-15% of acute human leukemias as part of a gene fusion with RUNX1 (also called AML1). Of the four ETO domains related to Drosophila melanogaster Nervy, only two are required to induce spontaneous myeloid leukemia upon transplantation into the mouse. One of these domains is related in sequence to TAF4, a component of TFIID. The structure of this domain, ETO-TAFH, is similar to yeast Rpb4 and to Escherichia coli sigma(70); it is the first TAF-related protein with structural similarity to the multisubunit RNA polymerases. Overlapping surfaces of ETO-TAFH interact with an autonomous repression domain of the nuclear receptor corepressor N-CoR and with a conserved activation domain from the E-box family of transcription factors. Thus, ETO-TAFH acts as a structural platform that can interchange negative and positive coregulatory proteins to control transcription.
Organizational Affiliation:
Laboratory of Molecular Biophysics, The Rockefeller University, 1230 York Avenue, New York, New York, 10021, USA.