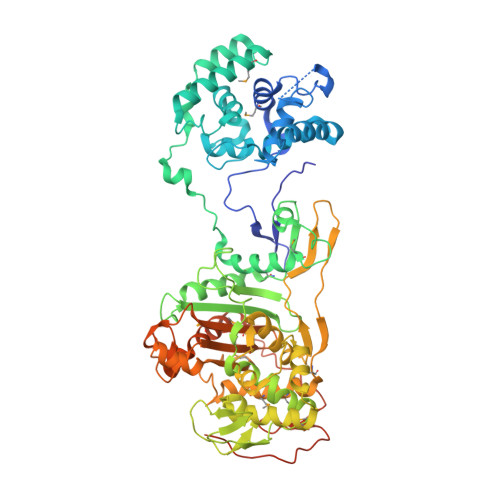
Structural insight into the transglycosylation step of bacterial cell-wall biosynthesis.
Lovering, A.L., de Castro, L.H., Lim, D., Strynadka, N.C.(2007) Science 315: 1402-1405
- PubMed: 17347437
- DOI: https://doi.org/10.1126/science.1136611
- Primary Citation of Related Structures:
2OLU, 2OLV - PubMed Abstract:
Peptidoglycan glycosyltransferases (GTs) catalyze the polymerization step of cell-wall biosynthesis, are membrane-bound, and are highly conserved across all bacteria. Long considered the "holy grail" of antibiotic research, they represent an essential and easily accessible drug target for antibiotic-resistant bacteria, including methicillin-resistant Staphylococcus aureus. We have determined the 2.8 angstrom structure of a bifunctional cell-wall cross-linking enzyme, including its transpeptidase and GT domains, both unliganded and complexed with the substrate analog moenomycin. The peptidoglycan GTs adopt a fold distinct from those of other GT classes. The structures give insight into critical features of the catalytic mechanism and key interactions required for enzyme inhibition.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, and Center for Blood Research, University of British Columbia, 2350 Health Sciences Mall, Vancouver, Canada.