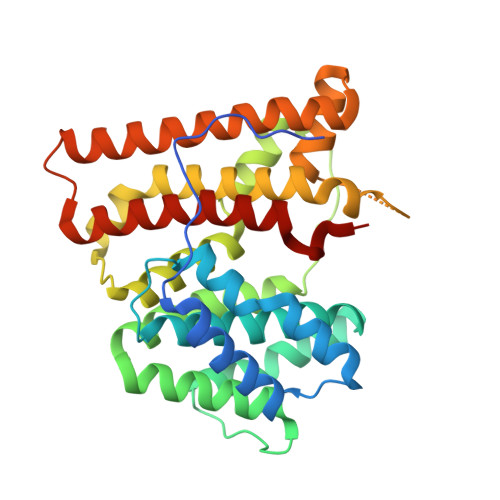
X-ray crystal structure of aristolochene synthase from Aspergillus terreus and evolution of templates for the cyclization of farnesyl diphosphate
Shishova, E.Y., Di Costanzo, L., Cane, D.E., Christianson, D.W.(2007) Biochemistry 46: 1941-1951
- PubMed: 17261032
- DOI: https://doi.org/10.1021/bi0622524
- Primary Citation of Related Structures:
2E4O, 2OA6 - PubMed Abstract:
Aristolochene synthase from Aspergillus terreus catalyzes the cyclization of the universal sesquiterpene precursor, farnesyl diphosphate, to form the bicyclic hydrocarbon aristolochene. The 2.2 A resolution X-ray crystal structure of aristolochene synthase reveals a tetrameric quaternary structure in which each subunit adopts the alpha-helical class I terpene synthase fold with the active site in the "open", solvent-exposed conformation. Intriguingly, the 2.15 A resolution crystal structure of the complex with Mg2+3-pyrophosphate reveals ligand binding only to tetramer subunit D, which is stabilized in the "closed" conformation required for catalysis. Tetramer assembly may hinder conformational changes required for the transition from the inactive open conformation to the active closed conformation, thereby accounting for the attenuation of catalytic activity with an increase in enzyme concentration. In both conformations, but especially in the closed conformation, the active site contour is highly complementary in shape to that of aristolochene, and a catalytic function is proposed for the pyrophosphate anion based on its orientation with regard to the presumed binding mode of aristolochene. A similar active site contour is conserved in aristolochene synthase from Penicillium roqueforti despite the substantial divergent evolution of these two enzymes, while strikingly different active site contours are found in the sesquiterpene cyclases 5-epi-aristolochene synthase and trichodiene synthase. Thus, the terpenoid cyclase active site plays a critical role as a template in binding the flexible polyisoprenoid substrate in the proper conformation for catalysis. Across the greater family of terpenoid cyclases, this template is highly evolvable within a conserved alpha-helical fold for the synthesis of terpene natural products of diverse structure and stereochemistry.
Organizational Affiliation:
Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, Pennsylvania 19104-6323, USA.