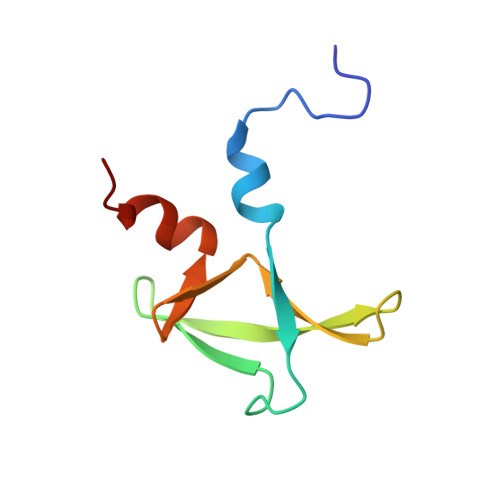
The C terminus of Pcf11 forms a novel zinc-finger structure that plays an essential role in mRNA 3'-end processing.
Yang, F., Hsu, P., Lee, S.D., Yang, W., Hoskinson, D., Xu, W., Moore, C., Varani, G.(2017) RNA 23: 98-107
- PubMed: 27780845
- DOI: https://doi.org/10.1261/rna.058354.116
- Primary Citation of Related Structures:
2NAX - PubMed Abstract:
3'-End processing of pre-mRNAs prior to packaging and export to the cytoplasm of the mature transcript is a highly regulated process executed by several tens of protein factors that recognize poorly conserved RNA signals. Among them is Pcf11, a highly conserved, multidomain protein that links transcriptional elongation, 3'-end processing, and transcription termination. Here we report the structure and biochemical function of Pcf11's C-terminal domain, which is conserved from yeast to humans. We identify a novel zinc-finger fold, resembling a trillium flower. Structural, biochemical, and genetic analyses reveal a highly conserved surface that plays a critical role in both cleavage and polyadenylation. These findings provide further insight into this important protein and its multiple functional roles during cotranscriptional RNA processing.
Organizational Affiliation:
Department of Chemistry, University of Washington, Seattle, Washington 98195, USA.