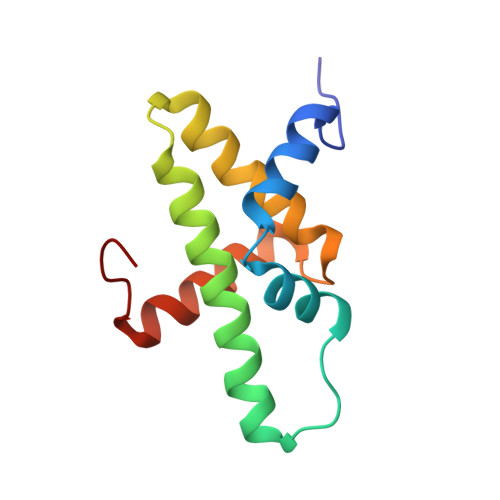
The Chromatin Remodelling Protein CHD1 Contains a Previously Unrecognised C-Terminal Helical Domain.
Mohanty, B., Helder, S., Silva, A.P., Mackay, J.P., Ryan, D.P.(2016) J Mol Biol 428: 4298-4314
- PubMed: 27591891
- DOI: https://doi.org/10.1016/j.jmb.2016.08.028
- Primary Citation of Related Structures:
2N39 - PubMed Abstract:
The packaging of eukaryotic DNA into nucleosomes, and the organisation of these nucleosomes into chromatin, plays a critical role in regulating all DNA-associated processes. Chromodomain helicase DNA-binding protein 1 (CHD1) is an ATP-dependent chromatin remodelling protein that is conserved throughout eukaryotes and has an ability to assemble and organise nucleosomes both in vitro and in vivo. This activity is involved in the regulation of transcription and is implicated in mammalian development and stem cell biology. CHD1 is classically depicted as possessing a pair of tandem chromodomains that directly precede a core catalytic helicase-like domain that is then followed by a SANT-SLIDE DNA-binding domain. Here, we have identified an additional conserved domain C-terminal to the SANT-SLIDE domain and determined its structure by multidimensional heteronuclear NMR spectroscopy. We have termed this domain the CHD1 helical C-terminal (CHCT) domain as it is comprised of five α-helices arranged in a variant helical bundle topology. CHCT has a conserved, positively charged surface and is able to bind DNA and nucleosomes. In addition, we have identified another group of proteins, the as yet uncharacterised C17orf64 proteins, as also containing a conserved CHCT domain. Our data provide new structural insights into the CHD1 enzyme family.
Organizational Affiliation:
School of Life and Environmental Sciences, The University of Sydney, Building G08, Corner Butlin Avenue and Maze Crescent, Sydney, New South Wales, 2006, Australia; Faculty of Pharmacy and Pharmaceutical Sciences, Medicinal Chemistry, Monash Institute of Pharmaceutical Sciences, Monash University, 381 Royal Parade, Parkville, Victoria, 3052, Australia.