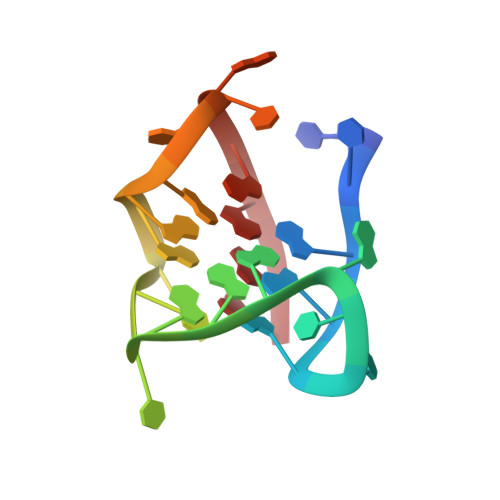
Solution structure of an intramolecular (3 + 1) human telomeric g-quadruplex bound to a telomestatin derivative.
Chung, W.J., Heddi, B., Tera, M., Iida, K., Nagasawa, K., Phan, A.T.(2013) J Am Chem Soc 135: 13495-13501
- PubMed: 23909929
- DOI: https://doi.org/10.1021/ja405843r
- Primary Citation of Related Structures:
2MB3 - PubMed Abstract:
Guanine-rich human telomeric DNA can adopt secondary structures known as G-quadruplexes, which can be targeted by small molecules to achieve anticancer effects. So far, the structural information on complexes between human telomeric DNA and ligands is limited to the parallel G-quadruplex conformation, despite the high structural polymorphism of human telomeric G-quadruplexes. No structure has been yet resolved for the complex with telomestatin, one of the most promising G-quadruplex-targeting anticancer drug candidates. Here we present the first high-resolution structure of the complex between an intramolecular (3 + 1) human telomeric G-quadruplex and a telomestatin derivative, the macrocyclic hexaoxazole L2H2-6M(2)OTD. This compound is observed to interact with the G-quadruplex through π-stacking and electrostatic interactions. This structural information provides a platform for the design of topology-specific G-quadruplex-targeting compounds and is valuable for the development of new potent anticancer drugs.
Organizational Affiliation:
School of Physical and Mathematical Sciences, Nanyang Technological University , Singapore.