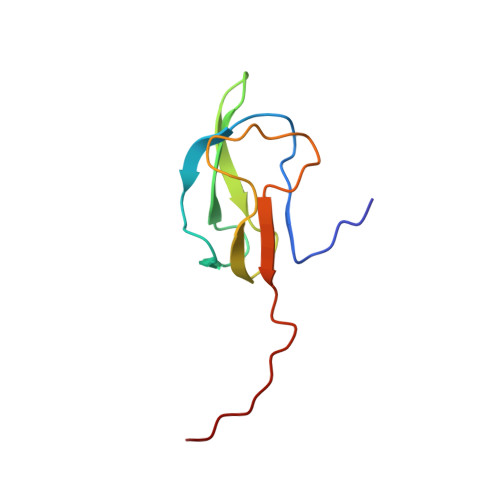
Biotinoyl domain of human acetyl-CoA carboxylase: Structural insights into the carboxyl transfer mechanism.
Lee, C.K., Cheong, H.K., Ryu, K.S., Lee, J.I., Lee, W., Jeon, Y.H., Cheong, C.(2008) Proteins 72: 613-624
- PubMed: 18247344
- DOI: https://doi.org/10.1002/prot.21952
- Primary Citation of Related Structures:
2KCC - PubMed Abstract:
Acetyl-CoA carboxylase (ACC) catalyzes the first step in fatty acid biosynthesis: the synthesis of malonyl-CoA from acetyl-CoA. As essential regulators of fatty acid biosynthesis and metabolism, ACCs are regarded as therapeutic targets for the treatment of metabolic diseases such as obesity. In ACC, the biotinoyl domain performs a critical function by transferring an activated carboxyl group from the biotin carboxylase domain to the carboxyl transferase domain, followed by carboxyl transfer to malonyl-CoA. Despite the intensive research on this enzyme, only the bacterial and yeast ACC structures are currently available. To explore the mechanism of ACC holoenzyme function, we determined the structure of the biotinoyl domain of human ACC2 and analyzed its characteristics and interaction with the biotin ligase, BirA using NMR spectroscopy. The 3D structure of the hACC2 biotinoyl domain has a similar folding topology to the earlier determined domains from E. coli and P. shermanii. However, the local structures near the biotinylation sites have notable differences that include the geometry of the consensus "Met-Lys-Met" (MKM) motif and the absence of "thumb" structure in the hACC2 biotinoyl domain. Observations of the NMR signals upon the biotinylation indicate that the biotin group of hACC2 does not affect the structure of the biotinoyl domain, while the biotin group for E. coli ACC interacts directly with the thumb residues that are not present in the hACC2 structure. These results imply that, in the E. coli ACC reaction, the biotin moiety carrying the carboxyl group from BC to CT can pause at the thumb of the BCCP domain. The human biotinoyl domain, however, lacks the thumb structure and does not have additional noncovalent interactions with the biotin moiety; thus, the flexible motion of the biotinylated lysine residue must underlie the "swinging arm" motion. The chemical shift perturbation and the cross saturation experiments of the human ACC2 holo-biotinoyl upon the addition of the biotin ligase (BirA) showed the interaction surface near the MKM motif, the two glutamic acids (Glu 926, Glu 953), and the positively charged residues (several lysine and arginine residues). This study provides insight into the mechanism of ACC holoenzyme function and supports the swinging arm model in human ACCs.
Organizational Affiliation:
The Magnetic Resonance Team, Korea Basic Science Institute, Ochang, Chungbuk 363-883, Korea.