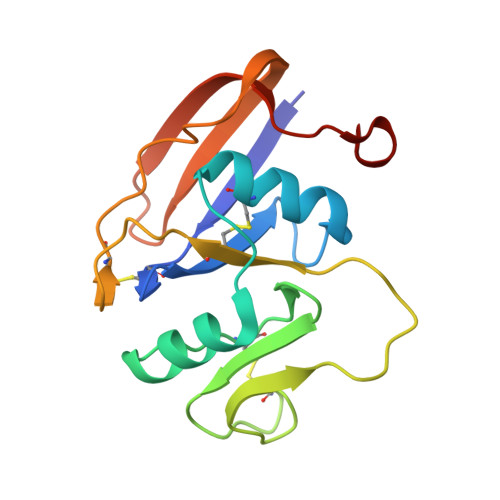
Structures of the Cd44-Hyaluronan Complex Provide Insight Into a Fundamental Carbohydrate-Protein Interaction.
Banerji, S., Wright, A.J., Noble, M.E.M., Mahoney, D.J., Campbell, I.D., Day, A.J., Jackson, D.G.(2008) Nat Struct Mol Biol 14: 234
- PubMed: 17293874
- DOI: https://doi.org/10.1038/nsmb1201
- Primary Citation of Related Structures:
2JCP, 2JCQ, 2JCR - PubMed Abstract:
Regulation of transient interactions between cells and the ubiquitous matrix glycosaminoglycan hyaluronan is crucial to such fundamental processes as embryonic development and leukocyte homing. Cd44, the primary cell surface receptor for hyaluronan, binds ligand via a lectin-like fold termed the Link module, but only after appropriate functional activation. The molecular details of the Cd44-hyaluronan interaction and hence the structural basis for this activation are unknown. Here we present the first crystal structure of Cd44 complexed with hyaluronan. This reveals that the interaction with hyaluronan is dominated by shape and hydrogen-bonding complementarity and identifies two conformational forms of the receptor that differ in orientation of a crucial hyaluronan-binding residue (Arg45, equivalent to Arg41 in human CD44). Measurements by NMR indicate that the conformational transition can be induced by hyaluronan binding, providing further insight into possible mechanisms for regulation of Cd44.
Organizational Affiliation:
Medical Research Council Human Immunology Unit, Weatherall Institute of Molecular Medicine, John Radcliffe Hospital, Headington, Oxford OX3 9DS, UK.