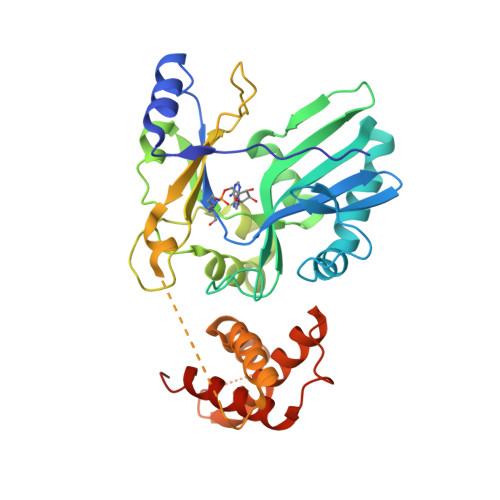
RNA Ligase Structures Reveal the Basis for RNA Specificity and Conformational Changes that Drive Ligation Forward.
Nandakumar, J., Shuman, S., Lima, C.D.(2006) Cell 127: 71-84
- PubMed: 17018278
- DOI: https://doi.org/10.1016/j.cell.2006.08.038
- Primary Citation of Related Structures:
2HVQ, 2HVR, 2HVS - PubMed Abstract:
T4 RNA ligase 2 (Rnl2) and kinetoplastid RNA editing ligases exemplify a family of RNA repair enzymes that seal 3'OH/5'PO(4) nicks in duplex RNAs via ligase adenylylation (step 1), AMP transfer to the nick 5'PO(4) (step 2), and attack by the nick 3'OH on the 5'-adenylylated strand to form a phosphodiester (step 3). Crystal structures are reported for Rnl2 at discrete steps along this pathway: the covalent Rnl2-AMP intermediate; Rnl2 bound to an adenylylated nicked duplex, captured immediately following step 2; and Rnl2 at an adenylylated nick in a state poised for step 3. These structures illuminate the stereochemistry of nucleotidyl transfer and reveal how remodeling of active-site contacts and conformational changes propel the ligation reaction forward. Mutational analysis and comparison of nick-bound structures of Rnl2 and human DNA ligase I highlight common and divergent themes of substrate recognition that can explain their specialization for RNA versus DNA repair.
Organizational Affiliation:
Structural Biology Program, Sloan-Kettering Institute, New York, NY 10021, USA.