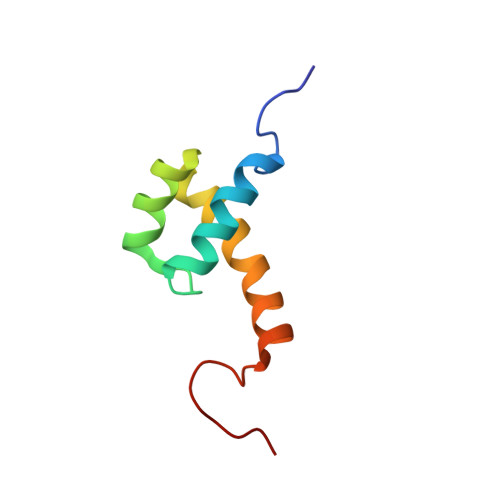
Analysis of the structure and function of the transcriptional coregulator HOP
Kook, H., Yung, W.W., Simpson, R.J., Kee, H.J., Shin, S., Lowry, J.A., Loughlin, F.E., Yin, Z., Epstein, J.A., Mackay, J.P.(2006) Biochemistry 45: 10584-10590
- PubMed: 16939210
- DOI: https://doi.org/10.1021/bi060641s
- Primary Citation of Related Structures:
2HI3 - PubMed Abstract:
Homeodomain-only protein (HOP) is an 8-kDa transcriptional corepressor that is essential for the normal development of the mammalian heart. Previous studies have shown that HOP, which consists entirely of a putative homeodomain, acts downstream of Nkx2.5 and associates with the serum response factor (SRF), repressing transcription from SRF-responsive genes. HOP is also able to recruit histone deacetylase (HDAC) activity, consistent with its ability to repress transcription. Unlike other classic homeodomain proteins, HOP does not appear to interact with DNA, although it has been unclear if this is because of an overall divergent structure or because of specific amino acid differences between HOP and other homeodomains. To work toward an understanding of HOP function, we have determined the 3D structure of full-length HOP and used a range of biochemical assays to define the parts of the protein that are functionally important for its repression activity. We show that HOP forms a classical homeodomain fold but that it cannot recognize double stranded DNA, a result that emphasizes the importance of caution in predicting protein function from sequence homology alone. We also demonstrate that two distinct regions on the surface of HOP are required for its ability to repress an SRF-driven reporter gene, and it is likely that these motifs direct interactions between HOP and partner proteins such as SRF- and HDAC-containing complexes. Our results demonstrate that the homeodomain fold has been co-opted during evolution for functions other than sequence-specific DNA binding and suggest that HOP functions as an adaptor protein to mediate transcriptional repression.
Organizational Affiliation:
Medical Research Center for Gene Regulation, Chonnam National University Medical School, Gwangju, 501-746, South Korea.