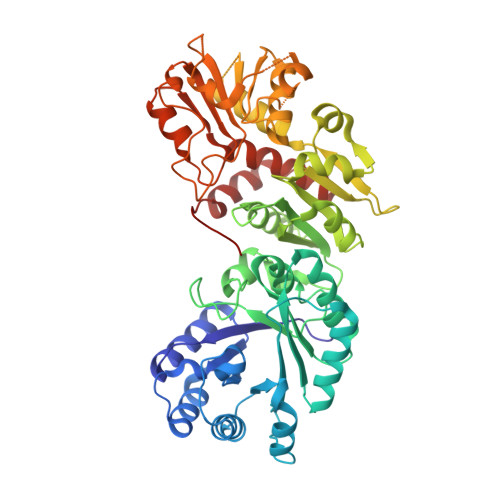
Structure of Arabidopsis dehydroquinate dehydratase-shikimate dehydrogenase and implications for metabolic channeling in the shikimate pathway
Singh, S.A., Christendat, D.(2006) Biochemistry 45: 7787-7796
- PubMed: 16784230
- DOI: https://doi.org/10.1021/bi060366+
- Primary Citation of Related Structures:
2GPT - PubMed Abstract:
The bifunctional enzyme dehydroquinate dehydratase-shikimate dehydrogenase (DHQ-SDH) catalyzes the dehydration of dehydroquinate to dehydroshikimate and the reduction of dehydroshikimate to shikimate in the shikimate pathway. We report the first crystal structure of Arabidopsis DHQ-SDH with shikimate bound at the SDH site and tartrate at the DHQ site. The interactions observed in the DHQ-tartrate complex reveal a conserved mode for substrate binding between the plant and microbial DHQ dehydratase family of enzymes. The SDH-shikimate complex provides the first direct evidence of the role of active site residues in the catalytic mechanism. Site-directed mutagenesis and mechanistic analysis revealed that Asp 423 and Lys 385 are key catalytic groups and Ser 336 is a key binding group. The arrangement of the two functional domains reveals that the control of metabolic flux through the shikimate pathway is achieved by increasing the effective concentration of dehydroshikimate through the proximity of the two sites.
Organizational Affiliation:
Department of Botany, University of Toronto, 25 Willcocks Street, Toronto, ON M5S 3B2, Canada.