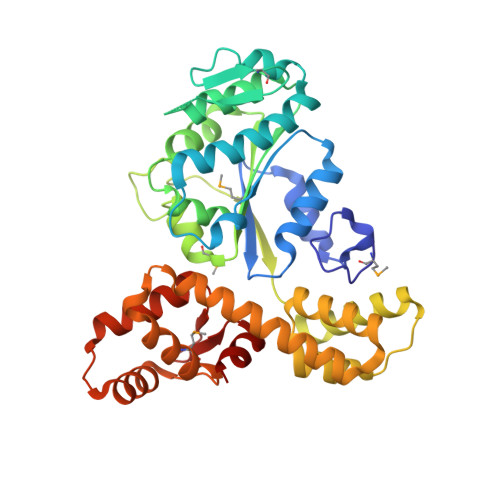
Crystal structure of a novel archaeal AAA+ ATPase SSO1545 from Sulfolobus solfataricus.
Xu, Q., Rife, C.L., Carlton, D., Miller, M.D., Krishna, S.S., Elsliger, M.A., Abdubek, P., Astakhova, T., Chiu, H.J., Clayton, T., Duan, L., Feuerhelm, J., Grzechnik, S.K., Hale, J., Han, G.W., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kumar, A., McMullan, D., Morse, A.T., Nigoghossian, E., Okach, L., Oommachen, S., Paulsen, J., Reyes, R., van den Bedem, H., Hodgson, K.O., Wooley, J., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2009) Proteins 74: 1041-1049
- PubMed: 19089981
- DOI: https://doi.org/10.1002/prot.22325
- Primary Citation of Related Structures:
2FNA
Organizational Affiliation:
Joint Center for Structural Genomics, Stanford Synchrotron Radiation Lightsource, Stanford University, Menlo Park, California, USA.