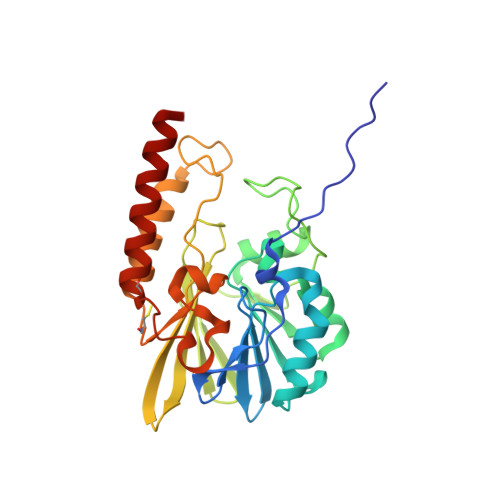
Structural insights into the design of inhibitors for the L1 metallo-beta-lactamase from Stenotrophomonas maltophilia.
Nauton, L., Kahn, R., Garau, G., Hernandez, J.F., Dideberg, O.(2008) J Mol Biol 375: 257-269
- PubMed: 17999929
- DOI: https://doi.org/10.1016/j.jmb.2007.10.036
- Primary Citation of Related Structures:
2FM6, 2FU7, 2FU8, 2FU9, 2GFJ, 2GFK, 2H6A, 2HB9 - PubMed Abstract:
One mechanism by which bacteria can escape the action of beta-lactam antibiotics is the production of metallo-beta-lactamases. Inhibition of these enzymes should restore the action of these widely used antibiotics. The tetrameric enzyme L1 from Stenotrophomonas maltophilia was used as a model system to determine a series of high-resolution crystal structures of apo, mono and bi-metal substituted proteins as well as protein-inhibitor complexes. Unexpectedly, although the apo structure revealed only few significant structural differences from the holo structure, some inhibitors were shown to induce amino acid side-chain rotations in the tightly packed active site. Moreover, one inhibitor employs a new binding mode in order to interact with the di-zinc center. This structural information could prove essential in the process of elucidation of the mode of interaction between a putative lead compound and metallo-beta-lactamases, one of the main steps in structure-based drug design.
Organizational Affiliation:
Institut de Biologie Structurale Jean-Pierre Ebel (UMR 5075;CNRS;CEA;UJF), 41, rue Jules Horowitz, F-38027 Grenoble Cedex 1, France.