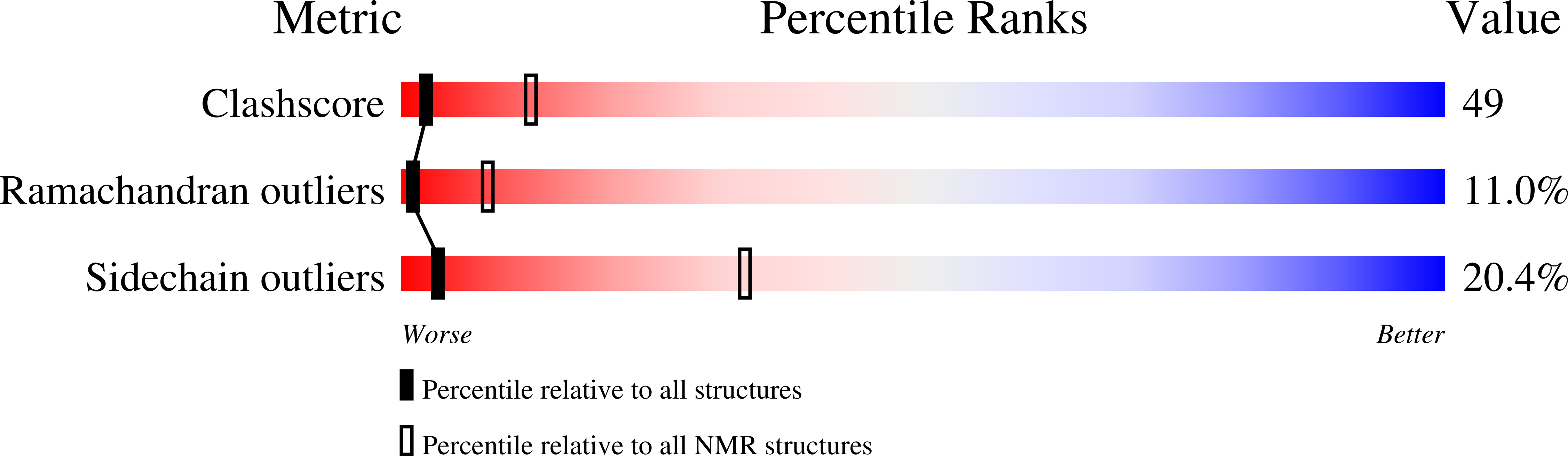Examination of the structure, stability, and catalytic potential in the engineered phosphoryl carrier domain of pyruvate phosphate dikinase.
Lin, Y., Lusin, J.D., Ye, D., Dunaway-Mariano, D., Ames, J.B.(2006) Biochemistry 45: 1702-1711
- PubMed: 16460017
- DOI: https://doi.org/10.1021/bi051816l
- Primary Citation of Related Structures:
2FM4 - PubMed Abstract:
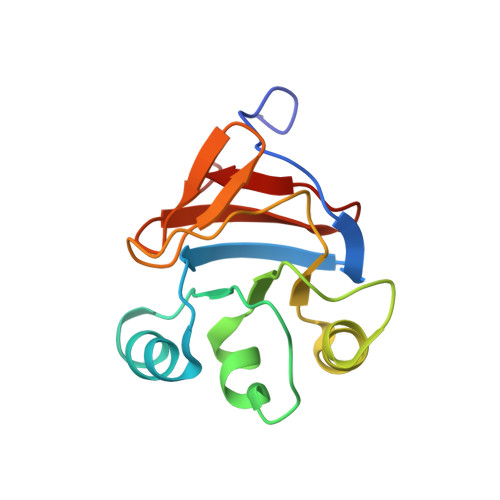
Pyruvate phosphate dikinase (PPDK) is a multidomain protein that catalyzes the interconversion of ATP, pyruvate, and phosphate with AMP, phosphoenolpyruvate (PEP), and pyrophosphate using its central domain to transport phosphoryl groups between two distant active sites. In this study, the mechanism by which the central domain moves between the two catalytic sites located on the N-terminal and C-terminal domains was probed by expressing this domain as an independent protein and measuring its structure, stability, and ability to catalyze the ATP/phosphate partial reaction in conjunction with the engineered N-terminal domain protein (residues 1-340 of the native PPDK). The encoding gene was engineered to express the central domain as residues 381-512 of the native PPDK. The central domain was purified and shown to be soluble, monomeric (13,438 Da), and stable (deltaG = 4.3 kcal/mol for unfolding in buffer at pH 7.0, 25 degrees C) and to possess native structure, as determined by multidimensional heteronuclear NMR analysis. The main chain structure of the central domain in solution aligns closely with that of the X-ray structure of native PPDK (the root-mean-square deviation is 2.2 A). Single turnover reactions of [14C]ATP and phosphate, carried out in the presence of equal concentrations of central domain and the N-terminal domain protein, did not produce the expected products, in contrast to efficient product formation observed for the N-terminal central domain construct (residues 1-553 of the native PPDK). These results are interpreted as evidence that the central domain, although solvent-compatible, must be tethered by the flexible linkers to the N-terminal domain for the productive domain-domain docking required for efficient catalysis.
Organizational Affiliation:
Department of Chemistry, University of New Mexico, Albuquerque, New Mexico 87131, USA.