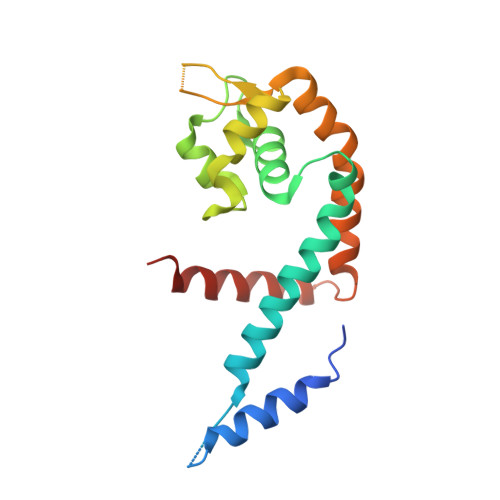
The Crystal Structure of the Transcriptional Regulator HucR from Deinococcus radiodurans Reveals a Repressor Preconfigured for DNA Binding.
Bordelon, T., Wilkinson, S.P., Grove, A., Newcomer, M.E.(2006) J Mol Biol 360: 168-177
- PubMed: 16750221
- DOI: https://doi.org/10.1016/j.jmb.2006.05.005
- Primary Citation of Related Structures:
2FBK - PubMed Abstract:
We report here the 2.3 A resolution structure of the hypothetical uricase regulator (HucR) from Deinococcus radiodurans R1. HucR, a member of the MarR family of DNA-binding proteins, was previously shown to repress its own expression as well as that of a uricase, a repression that is alleviated both in vivo and in vitro upon binding uric acid, the substrate for uricase. As uric acid is a potent scavenger of reactive oxygen species, and as D. radiodurans is known for its remarkable resistance to DNA-damaging agents, these observations indicate a novel oxidative stress response mechanism. The crystal structure of HucR in the absence of ligand or DNA reveals a dimer in which the DNA recognition helices are preconfigured for DNA binding. This configuration of DNA-binding domains is achieved through an apparently stable dimer interface that, in contrast to what is observed in other MarR homologs for which structures have been determined, shows little conformational heterogeneity in the absence of ligand. An additional amino-terminal segment, absent from other MarR homologs, appears to brace the principal helix of the dimerization interface. However, although HucR is preconfigured for DNA binding, the presence of a stacked pair of symmetry-related histidine residues at a central pivot point in the dimer interface suggests a mechanism for a conformational change to attenuate DNA binding.
Organizational Affiliation:
Department of Biological Sciences, Louisiana State University, Baton Rouge, LA 70803, USA.