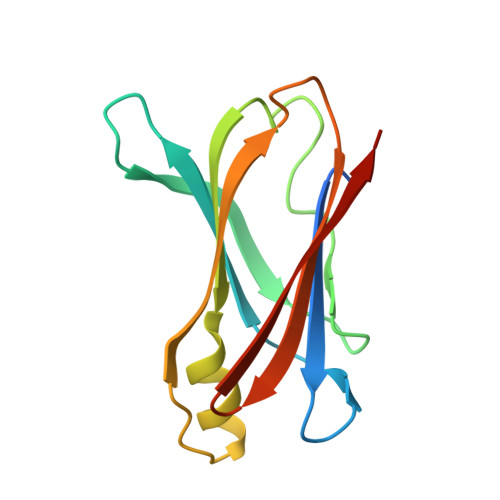
Diflunisal Analogues Stabilize the Native State of Transthyretin. Potent Inhibition of Amyloidogenesis.
Adamski-Werner, S.L., Palaninathan, S.K., Sacchettini, J.C., Kelly, J.W.(2004) J Med Chem 47: 355-374
- PubMed: 14711308
- DOI: https://doi.org/10.1021/jm030347n
- Primary Citation of Related Structures:
2B77, 2B9A, 2F7I, 3D2T - PubMed Abstract:
Analogues of diflunisal, an FDA-approved nonsteroidal antiinflammatory drug (NSAID), were synthesized and evaluated as inhibitors of transthyretin (TTR) aggregation, including amyloid fibril formation. High inhibitory activity was observed for 26 of the compounds. Of those, eight exhibited excellent binding selectivity for TTR in human plasma (binding stoichiometry >0.50, with a theoretical maximum of 2.0 inhibitors bound per TTR tetramer). Biophysical studies reveal that these eight inhibitors dramatically slow tetramer dissociation (the rate-determining step of amyloidogenesis) over a duration of 168 h. This appears to be achieved through ground-state stabilization, which raises the kinetic barrier for tetramer dissociation. Kinetic stabilization of WT TTR by these eight inhibitors is further substantiated by the decreasing rate of amyloid fibril formation as a function of increasing inhibitor concentration (pH 4.4). X-ray cocrystal structures of the TTR.18(2) and TTR.20(2) complexes reveal that 18 and 20 bind in opposite orientations in the TTR binding site. Moving the fluorines from the meta positions in 18 to the ortho positions in 20 reverses the binding orientation, allowing the hydrophilic aromatic ring of 20 to orient in the outer binding pocket where the carboxylate engages in favorable electrostatic interactions with the epsilon-ammonium groups of Lys 15 and 15'. The hydrophilic aryl ring of 18 occupies the inner binding pocket, with the carboxylate positioned to hydrogen bond to the serine 117 and 117' residues. Diflunisal itself appears to occupy both orientations based on the electron density in the TTR.1(2) structure. Structure-activity relationships reveal that para-carboxylate substitution on the hydrophilic ring and dihalogen substitution on the hydrophobic ring afford the most active TTR amyloid inhibitors.
Organizational Affiliation:
The Department of Chemistry and the Skaggs Institute of Chemical Biology, The Scripps Research Institiute, 10550 N. Torrey Pines Rd., La Jolla, California 92037, USA.