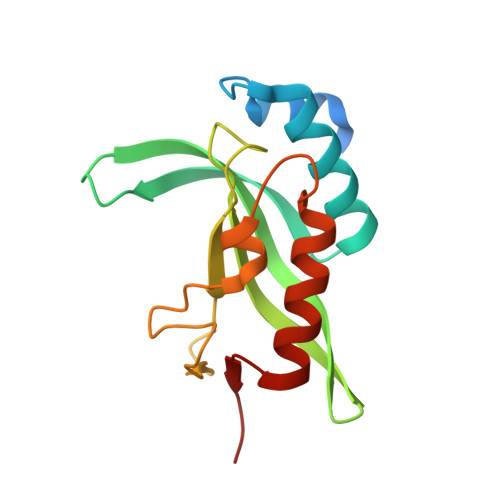
Structure of human TSG101 UEV domain.
Palencia, A., Martinez, J.C., Mateo, P.L., Luque, I., Camara-Artigas, A.(2006) Acta Crystallogr D Biol Crystallogr 62: 458-464
- PubMed: 16552148
- DOI: https://doi.org/10.1107/S0907444906005221
- Primary Citation of Related Structures:
2F0R - PubMed Abstract:
The UEV domain of the TSG101 protein functions in the vacuolar protein-sorting pathway and in the budding process of HIV-1 and other retroviruses by recognizing ubiquitin in proteins tagged for degradation and short sequences in viral proteins containing an essential and well conserved PTAP motif, respectively. A deep understanding of these interactions is key to the rational design of much-needed novel antivirals. Here, the crystal structure of the TSG101 UEV domain (TSG101-UEV) is presented. TSG101-UEV was crystallized in the presence of PEG 4000 and ammonium sulfate. Under these conditions, crystals were obtained in space group R3, with unit-cell parameters a = b = 97.9, c = 110.6 A, alpha = beta = 90, gamma = 120 degrees . Phases were solved by molecular replacement and the crystal structure of TSG101-UEV was refined to an R factor of 18.8% at 2.2 A resolution. A comparison between the crystal structure and previously reported NMR structures has revealed significant differences in the conformation of one of the loops implicated in ubiquitin recognition. Also, the resulting structure has provided information about the presence of water molecules at the binding interface that could be of relevance for peptide recognition.
Organizational Affiliation:
Department of Physical Chemistry and Institute of Biotechnology, Faculty of Sciences, University of Granada, 18071 Granada, Spain.