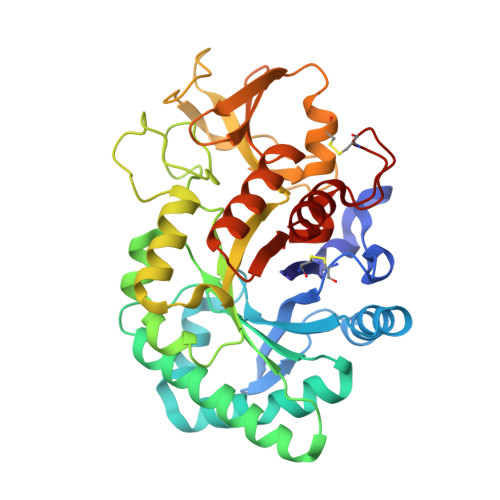
Structure of a bovine secretory signalling glycoprotein (SPC-40) at 2.1 Angstrom resolution.
Kumar, J., Ethayathulla, A.S., Srivastava, D.B., Sharma, S., Singh, S.B., Srinivasan, A., Yadav, M.P., Singh, T.P.(2006) Acta Crystallogr D Biol Crystallogr 62: 953-963
- PubMed: 16929095
- DOI: https://doi.org/10.1107/S0907444906020427
- Primary Citation of Related Structures:
2ESC - PubMed Abstract:
A recently discovered new class of 40 kDa glycoproteins forms a major component of the secretory proteins in the dry secretions of non-lactating animals. These proteins are implicated as protective signalling factors that determine which cells are to survive during the processes of drastic tissue remodelling. In order to understand its role in the remodelling of mammary glands, the detailed three-dimensional structure of the bovine signalling glycoprotein (SPC-40) has been determined using X-ray crystallography. SPC-40 was purified from bovine dry secretions and crystallized using the hanging-drop vapour-diffusion method. The crystals belong to the orthorhombic space group P2(1)2(1)2(1), with unit-cell parameters a = 62.6, b = 67.4, c = 106.9 Angstrom. The protein was also cloned in order to determine its complete amino-acid sequence. Its three-dimensional structure has been determined using data to 2.1 Angstrom resolution. The amino-acid sequence determination of SPC-40 reveals two potential N-glycosylation sites at Asn39 and Asn345, but electron density for a glycan chain was only present at Asn39. The protein adopts a conformation with the classical (beta/alpha)(8)-barrel fold of triosephosphate isomerase (TIM barrel; residues 1-237 and 310-360) with the insertion of a small alpha+beta domain (residues 240-307) similar to that observed in chitinases. However, the substitution of Leu for Glu in the consensus catalytic sequence in SPC-40 caused a loss of chitinase activity. Furthermore, the chitin-binding groove in SPC-40 is considerably distorted owing to unfavourable conformations of several residues, including Trp78, Tyr120, Asp186 and Arg242. Three surface loops, His188-His197, Phe202-Arg212 and Tyr244-Pro260, have exceptionally high B factors, suggesting large-scale flexibility. Fluorescence studies indicate that various sugars bind to SPC-40 with low affinities.
Organizational Affiliation:
Department of Biophysics, All India Institute of Medical Sciences, New Delhi 110029, India.