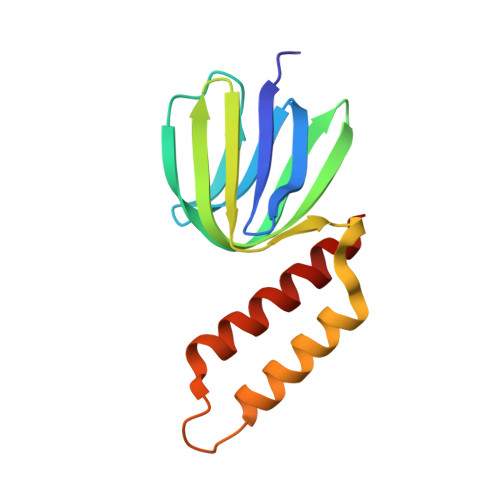
Structures of the thermophilic F1-ATPase {varepsilon} subunit suggesting ATP-regulated arm motion of its C-terminal domain in F1
Yagi, H., Kajiwara, N., Tanaka, H., Tsukihara, T., Kato-Yamada, Y., Yoshida, M., Akutsu, H.(2007) Proc Natl Acad Sci U S A 104: 11233-11238
- PubMed: 17581881
- DOI: https://doi.org/10.1073/pnas.0701045104
- Primary Citation of Related Structures:
2E5T, 2E5U, 2E5Y - PubMed Abstract:
The epsilon subunit of bacterial and chloroplast F(o)F(1)-ATP synthases modulates their ATP hydrolysis activity. Here, we report the crystal structure of the ATP-bound epsilon subunit from a thermophilic Bacillus PS3 at 1.9-A resolution. The C-terminal two alpha-helices were folded into a hairpin, sitting on the beta sandwich structure, as reported for Escherichia coli. A previously undescribed ATP binding motif, I(L)DXXRA, recognizes ATP together with three arginine and one glutamate residues. The E. coli epsilon subunit binds ATP in a similar manner, as judged on NMR. We also determined solution structures of the C-terminal domain of the PS3 epsilon subunit and relaxation parameters of the whole molecule by NMR. The two helices fold into a hairpin in the presence of ATP but extend in the absence of ATP. The latter structure has more helical regions and is much more flexible than the former. These results suggest that the epsilon C-terminal domain can undergo an arm-like motion in response to an ATP concentration change and thereby contribute to regulation of F(o)F(1)-ATP synthase.
Organizational Affiliation:
Institute for Protein Research, Osaka University, 3-2 Yamadaoka, Suita 565-0871, Japan.