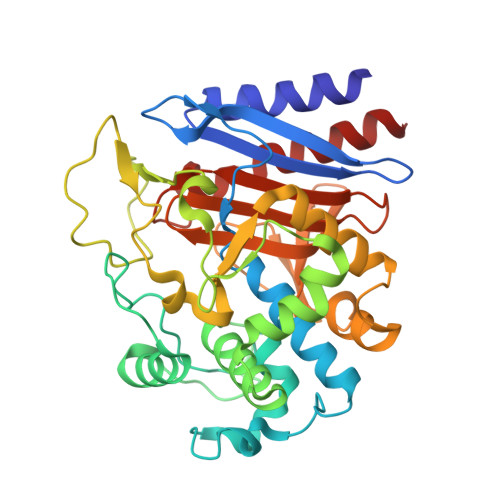
Crystal Structure and Functional Characterization of a D-Stereospecific Amino Acid Amidase from Ochrobactrum anthropi SV3, a New Member of the Penicillin-recognizing Proteins
Okazaki, S., Suzuki, A., Komeda, H., Yamaguchi, S., Asano, Y., Yamane, T.(2007) J Mol Biol 368: 79-91
- PubMed: 17331533
- DOI: https://doi.org/10.1016/j.jmb.2006.10.070
- Primary Citation of Related Structures:
2DNS, 2DRW - PubMed Abstract:
D-amino acid amidase (DAA) from Ochrobactrum anthropi SV3, which catalyzes the stereospecific hydrolysis of D-amino acid amides to yield the D-amino acid and ammonia, has attracted increasing attention as a catalyst for the stereospecific production of D-amino acids. In order to clarify the structure-function relationships of DAA, the crystal structures of native DAA, and of the D-phenylalanine/DAA complex, were determined at 2.1 and at 2.4 A resolution, respectively. Both crystals contain six subunits (A-F) in the asymmetric unit. The fold of DAA is similar to that of the penicillin-recognizing proteins, especially D-alanyl-D-alanine-carboxypeptidase from Streptomyces R61, and class C beta-lactamase from Enterobacter cloacae strain GC1. The catalytic residues of DAA and the nucleophilic water molecule for deacylation were assigned based on these structures. DAA has a flexible Omega-loop, similar to class C beta-lactamase. DAA forms a pseudo acyl-enzyme intermediate between Ser60 O(gamma) and the carbonyl moiety of d-phenylalanine in subunits A, B, C, D, and E, but not in subunit F. The difference between subunit F and the other subunits (A, B, C, D and E) might be attributed to the order/disorder structure of the Omega-loop: the structure of this loop cannot assigned in subunit F. Deacylation of subunit F may be facilitated by the relative movement of deprotonated His307 toward Tyr149. His307 N(epsilon2) extracts the proton from Tyr149 O(eta), then Tyr149 O(eta) attacks a nucleophilic water molecule as a general base. Gln214 on the Omega-loop is essential for forming a network of water molecules that contains the nucleophilic water needed for deacylation. Although peptidase activity is found in almost all penicillin-recognizing proteins, DAA lacks peptidase activity. The lack of transpeptidase and carboxypeptidase activities may be attributed to steric hindrance of the substrate-binding pocket by a loop comprised of residues 278-290 and the Omega-loop.
Organizational Affiliation:
Department of Biotechnology, School of Engineering, Nagoya University, Chikusa, Nagoya 464-8603, Japan.