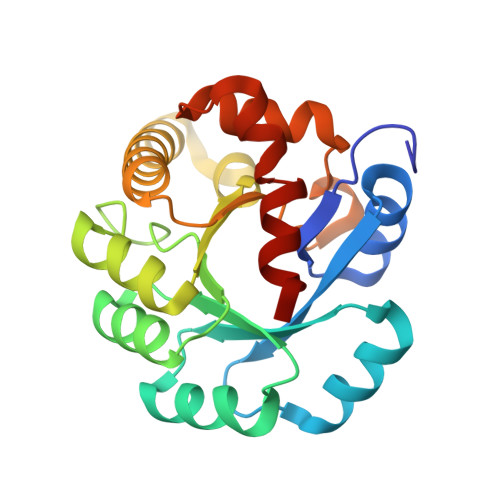
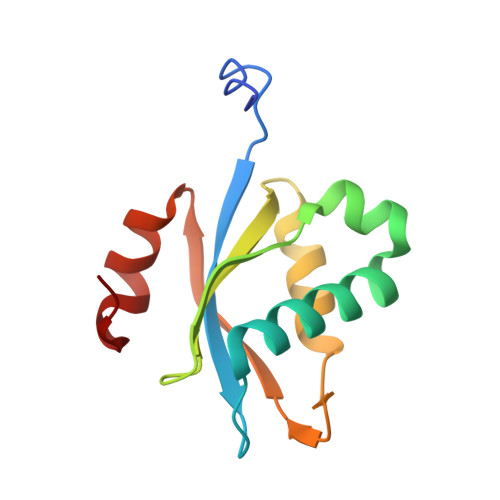
Crystal structure of protein Ph1481p in complex with protein Ph1877p of archaeal RNase P from Pyrococcus horikoshii OT3: implication of dimer formation of the holoenzyme
Kawano, S., Nakashima, T., Kakuta, Y., Tanaka, I., Kimura, M.(2006) J Mol Biol 357: 583-591
- PubMed: 16430919
- DOI: https://doi.org/10.1016/j.jmb.2005.12.086
- Primary Citation of Related Structures:
2CZV - PubMed Abstract:
Ribonuclease P (RNase P) in the hyperthermophilic archaeon Pyrococcus horikoshii OT3 consists of a catalytic RNA and five protein subunits. We previously determined crystal structures of four protein subunits. Ph1481p, an archaeal homologue for human hPop5, is the protein component of the P.horikoshii RNase P for which no structural information is available. Here we report the crystal structure of Ph1481p in complex with another protein subunit, Ph1877p, determined at 2.0 A resolution. Ph1481p consists of a five-stranded antiparallel beta-sheet and five helices, which fold in a way that is topologically similar to the ribonucleoprotein (RNP) domain. Ph1481p is, however, distinct from the typical RNP domain in that it has additional helices at the C terminus, which pack against one face of the beta-sheet. The presence of two complexes in the asymmetric unit, together with gel filtration chromatography indicates that the heterotetramer is stable in solution and represents a fundamental building block in the crystals. In the heterotetrameric structure (Ph1877p-(Ph1481p)(2)-Ph1877p), a homodimer of Ph1481p sits between two Ph1877p monomers. Ph1481p dimerizes through hydrogen bonding interaction from the loop between alpha1 and alpha2 helices, and each Ph1481p interacts with two Ph1877p molecules, where alpha2 and alpha3 in Ph1481p interact with alpha7 in one Ph1877p and alpha8 in the other Ph1877p molecule, respectively. Deletion of the alpha1-alpha2 loop in Ph1481p caused heterodimerization with Ph1877p, and abolished ability to homodimerize itself and heterotetramerize with Ph1877p. Furthermore, the reconstituted particle containing the deletion mutant Ph1481p (mPh1481p) exhibited significantly reduced nuclease activity. These results suggest the presence of the heterotetramer of Ph1481p and Ph1877p in P.horikoshii RNase P.
Organizational Affiliation:
Laboratory of Biochemistry, Department of Bioscience and Biotechnology, Faculty of Agriculture, Graduate School, Kyushu University, Fukuoka 812-8581, Japan.