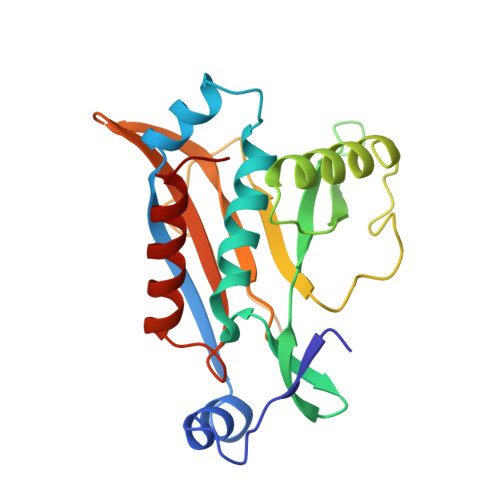
Crystal structure of the aspartic acid-199----asparagine mutant of chloramphenicol acetyltransferase to 2.35-A resolution: structural consequences of disruption of a buried salt bridge.
Gibbs, M.R., Moody, P.C., Leslie, A.G.(1990) Biochemistry 29: 11261-11265
- PubMed: 2271709
- DOI: https://doi.org/10.1021/bi00503a015
- Primary Citation of Related Structures:
2CLA - PubMed Abstract:
The crystal structure of the Asp-199----Asn mutant of chloramphenicol acetyltransferase (CAT) has been determined to 2.35-A resolution. In wild-type CAT Asp-199 is involved in a fully buried intrasubunit salt bridge with Arg-18, an interaction that is adjacent to the active site. Replacement of aspartate with asparagine by site-directed mutagenesis disrupts this salt bridge and causes extensive conformational changes within the active site. The imidazole group of the catalytically essential His-195 is reoriented, with the loss of interactions thought to stabilize the preferred tautomer of this residue. Arg-18 and Asn-199 form three new intersubunit interactions as a result of large side-chain torsion angle changes which cause the movement of two polypeptide loops, some residues of which are up to 20 A away from the site of the mutation. The new interactions of Arg-18 and Asn-199 compensate for the loss of the buried salt bridge and afford near-wild-type thermostability to Asn-199 CAT, albeit with a greatly reduced activity.
Organizational Affiliation:
Blackett Laboratory, Imperial College of Science, Technology, and Medicine, London, U.K.