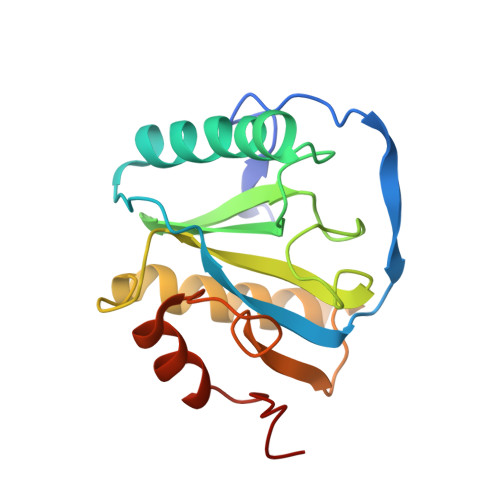
Structural Basis for Preferential Recognition of Diaminopimelic Acid-Type Peptidoglycan by a Subset of Peptidoglycan Recognition Proteins
Lim, J.-H., Kim, M.-S., Kim, H.-E., Yano, T., Oshima, Y., Aggarwal, K., Goldman, W.E., Silverman, N., Kurata, S., Oh, B.-H.(2006) J Biol Chem 281: 8286
- PubMed: 16428381
- DOI: https://doi.org/10.1074/jbc.M513030200
- Primary Citation of Related Structures:
2CB3 - PubMed Abstract:
Drosophila peptidoglycan recognition protein (PGRP)-LCx and -LCa are receptors that preferentially recognize meso-diaminopimelic acid (DAP)-type peptidoglycan (PGN) present in Gram-negative bacteria over lysine-type PGN of gram-positive bacteria and initiate the IMD signaling pathway, whereas PGRP-LE plays a synergistic role in this process of innate immune defense. How these receptors can distinguish the two types of PGN remains unclear. Here the structure of the PGRP domain of Drosophila PGRP-LE in complex with tracheal cytotoxin (TCT), the monomeric DAP-type PGN, reveals a buried ionic interaction between the unique carboxyl group of DAP and a previously unrecognized arginine residue. This arginine is conserved in the known DAP-type PGN-interacting PGRPs and contributes significantly to the affinity of the protein for the ligand. Unexpectedly, TCT induces infinite head-to-tail dimerization of PGRP-LE, in which the disaccharide moiety, but not the peptide stem, of TCT is positioned at the dimer interface. A sequence comparison suggests that TCT induces heterodimerization of the ectodomains of PGRP-LCx and -LCa in a closely analogous manner to prime the IMD signaling pathway, except that the heterodimer formation is nonperpetuating.
Organizational Affiliation:
Center for Biomolecular Recognition and Division of Molecular and Life Science, Department of Life Science, Pohang University of Science and Technology, Pohang, Kyungbuk 790-784, Korea.