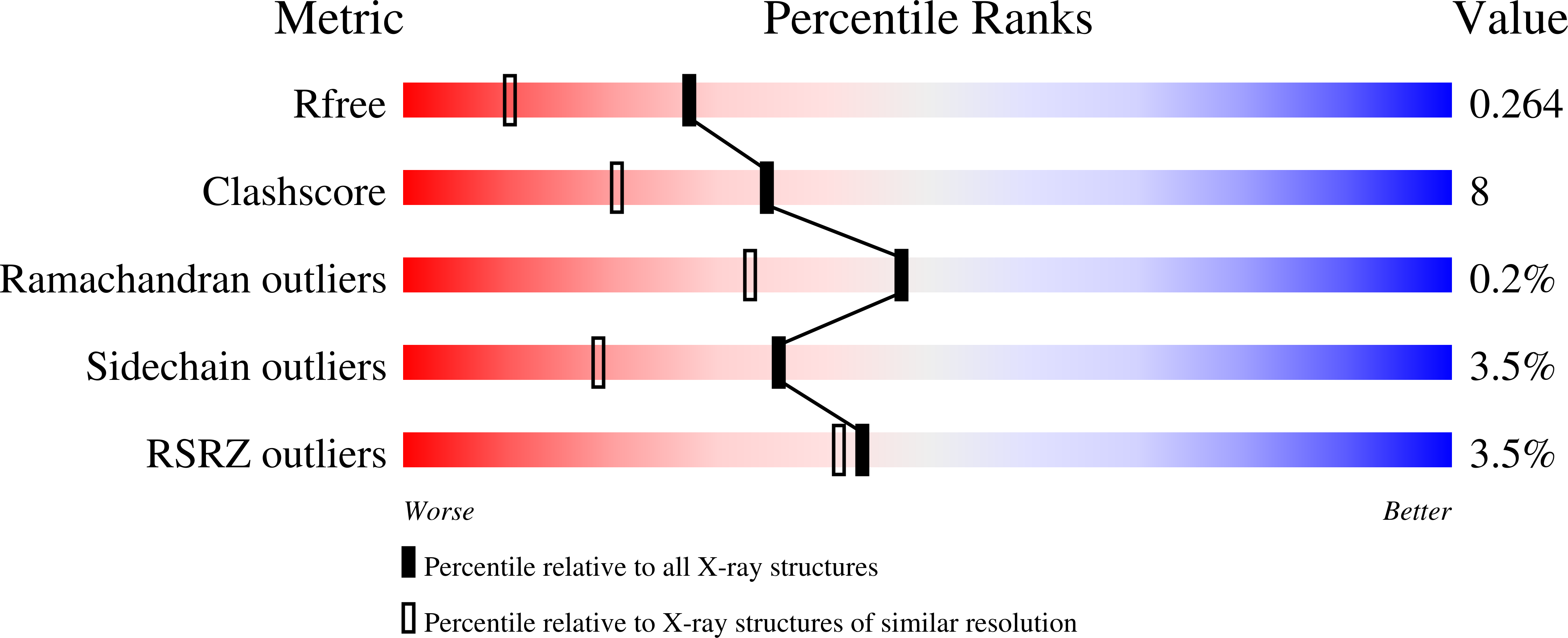
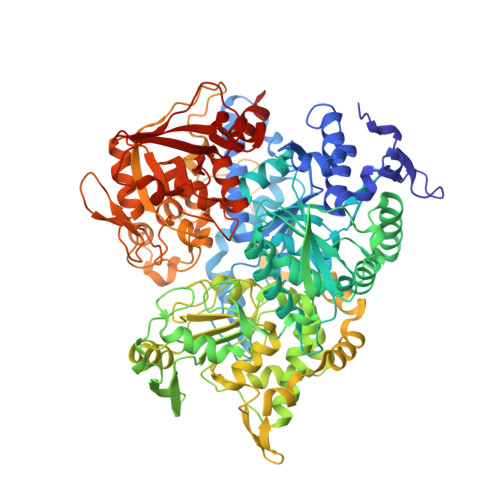
Crystal structure of human iron regulatory protein 1 as cytosolic aconitase
Dupuy, J., Volbeda, A., Carpentier, P., Darnault, C., Moulis, J.M., Fontecilla-Camps, J.C.(2006) Structure 14: 129-139
- PubMed: 16407072
- DOI: https://doi.org/10.1016/j.str.2005.09.009
- Primary Citation of Related Structures:
2B3X, 2B3Y - PubMed Abstract:
Iron regulatory proteins (IRPs) control the translation of proteins involved in iron uptake, storage and utilization by binding to specific noncoding sequences of the corresponding mRNAs known as iron-responsive elements (IREs). This strong interaction assures proper iron homeostasis in animal cells under iron shortage. Conversely, under iron-replete conditions, IRP1 binds a [4Fe-4S] cluster and functions as cytosolic aconitase. Regulation of the balance between the two IRP1 activities is complex, and it does not depend only on iron availability. Here, we report the crystal structure of human IRP1 in its aconitase form. Comparison with known structures of homologous enzymes reveals well-conserved folds and active site environments with significantly different surface shapes and charge distributions. The specific features of human IRP1 allow us to propose a tentative model of an IRP1-IRE complex that agrees with a range of previously obtained data.
Organizational Affiliation:
Laboratoire de Cristallographie et de Cristallogenèse des Protéines, Institut de Biologie Structurale JP Ebel, CEA/CNRS/Université Joseph Fourier, 38027 Grenoble Cedex 1, France.