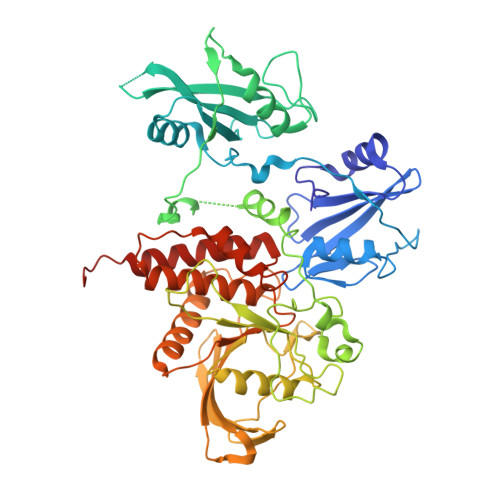
Crystal structure of human protein-tyrosine phosphatase SHP-1.
Yang, J., Liu, L., He, D., Song, X., Liang, X., Zhao, Z.J., Zhou, G.W.(2003) J Biol Chem 278: 6516-6520
- PubMed: 12482860
- DOI: https://doi.org/10.1074/jbc.M210430200
- PubMed Abstract:
SHP-1 is a cytosolic protein-tyrosine phosphatase that behaves as a negative regulator in eukaryotic cellular signaling pathways. To understand its regulatory mechanism, we have determined the crystal structure of the C-terminal truncated human SHP-1 in the inactive conformation at 2.8-A resolution and refined the structure to a crystallographic R-factor of 24.0%. The three-dimensional structure shows that the ligand-free SHP-1 has an auto-inhibited conformation. Its N-SH2 domain blocks the catalytic domain and keeps the enzyme in the inactive conformation, which supports that the phosphatase activity of SHP-1 is primarily regulated by the N-SH2 domain. In addition, the C-SH2 domain of SHP-1 has a different orientation from and is more flexible than that of SHP-2, which enables us to propose an enzymatic activation mechanism in which the C-SH2 domains of SHPs could be involved in searching for phosphotyrosine activators.
Organizational Affiliation:
Program in Molecular Medicine, University of Massachusetts Medical School, Worcester, Massachusetts 01605, USA.