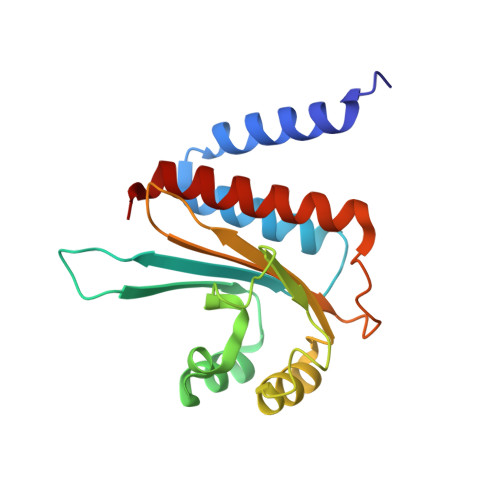
Structure of the Escherichia coli quorum sensing protein SdiA: activation of the folding switch by acyl homoserine lactones.
Yao, Y., Martinez-Yamout, M.A., Dickerson, T.J., Brogan, A.P., Wright, P.E., Dyson, H.J.(2006) J Mol Biol 355: 262-273
- PubMed: 16307757
- DOI: https://doi.org/10.1016/j.jmb.2005.10.041
- Primary Citation of Related Structures:
2AVX - PubMed Abstract:
The three-dimensional structure of a complex between the N-terminal domain of the quorum sensing protein SdiA of Escherichia coli and a candidate autoinducer N-octanoyl-L-homoserine lactone (C8-HSL) has been calculated in solution from NMR data. The SdiA-HSL system shows the "folding switch" behavior that has been seen for quorum-sensing factors produced by other bacterial species. In the presence of C8-HSL, a significant proportion of the SdiA protein is produced in a folded, soluble form in an E.coli expression system, whereas in the absence of acyl homoserine lactones, the protein is expressed into insoluble inclusion bodies. In the three-dimensional structure, the autoinducer molecule is sequestered in a deep pocket in the hydrophobic core, forming an integral part of the core packing of the folded SdiA. The NMR spectra of the complex show that the bound C8-HSL is conformationally heterogeneous, either due to motion within the pocket or to heterogeneity of the bound structure. The C8-HSL conformation is defined by NOEs to the protein only at the terminal methyl group of the octanoyl chain. Unlike other well-studied bacterial quorum sensing systems such as LuxR of Vibrio fischeri and TraR of Agrobacterium tumefaciens, there is no endogenous autoinducer for SdiA in E.coli: the E.coli genome does not contain a gene analogous to the LuxI and TraI autoinducer synthetases. We show that two other homoserine lactone derivatives are also capable of acting as a folding-switch autoinducers for SdiA. The observed structural heterogeneity of the bound C8-HSL in the complex, together with the variety of autoinducer-type molecules that can apparently act as folding switches in this system, are consistent with the postulated biological function of the SdiA protein as a detector of the presence of other species of bacteria.
Organizational Affiliation:
Department of Molecular Biology and the Skaggs Institute for Chemical Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.