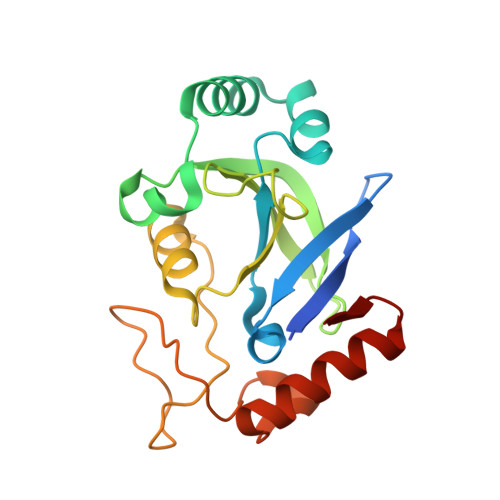
Crystal structures of the editing domain of Escherichia coli leucyl-tRNA synthetase and its complexes with Met and Ile reveal a lock-and-key mechanism for amino acid discrimination
Liu, Y., Liao, J., Zhu, B., Wang, E.D., Ding, J.(2006) Biochem J 394: 399-407
- PubMed: 16277600
- DOI: https://doi.org/10.1042/BJ20051249
- Primary Citation of Related Structures:
2AJG, 2AJH, 2AJI - PubMed Abstract:
aaRSs (aminoacyl-tRNA synthetases) are responsible for the covalent linking of amino acids to their cognate tRNAs via the aminoacylation reaction and play a vital role in maintaining the fidelity of protein synthesis. LeuRS (leucyl-tRNA synthetase) can link not only the cognate leucine but also the nearly cognate residues Ile and Met to tRNA(Leu). The editing domain of LeuRS deacylates the mischarged Ile-tRNA(Leu) and Met-tRNA(Leu). We report here the crystal structures of ecLeuRS-ED (the editing domain of Escherichia coli LeuRS) in both the apo form and in complexes with Met and Ile at 2.0 A, 2.4 A, and 3.2 A resolution respectively. The editing active site consists of a number of conserved amino acids, which are involved in the precise recognition and binding of the noncognate amino acids. The substrate-binding pocket has a rigid structure which has an optimal stereochemical fit for Ile and Met, but has steric hindrance for leucine. Based on our structural results and previously available biochemical data, we propose that ecLeuRS-ED uses a lock-and-key mechanism to recognize and discriminate between the amino acids. Structural comparison also reveals that all subclass Ia aaRSs share a conserved structure core consisting of the editing domain and conserved residues at the editing active site, suggesting that these enzymes may use a common mechanism for the editing function.
Organizational Affiliation:
Key Laboratory of Proteomics, Institute of Biochemistry and Cell Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, 320 Yue-Yang Road, Shanghai 200031, China.