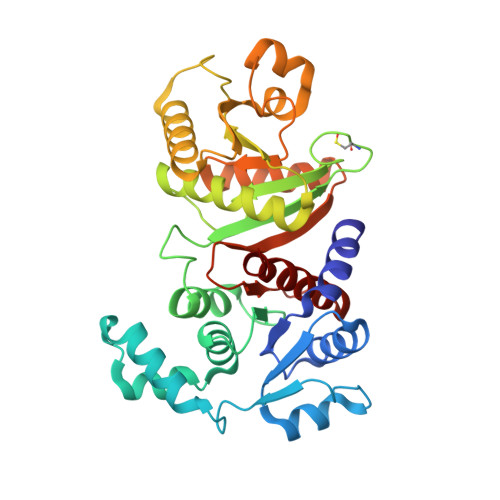
Structural and functional studies suggest a catalytic mechanism for the phosphotransacetylase from Methanosarcina thermophila.
Lawrence, S.H., Luther, K.B., Schindelin, H., Ferry, J.G.(2006) J Bacteriol 188: 1143-1154
- PubMed: 16428418
- DOI: https://doi.org/10.1128/JB.188.3.1143-1154.2006
- Primary Citation of Related Structures:
2AF3, 2AF4 - PubMed Abstract:
Phosphotransacetylase (EC 2.3.1.8) catalyzes reversible transfer of the acetyl group from acetyl phosphate to coenzyme A (CoA), forming acetyl-CoA and inorganic phosphate. Two crystal structures of phosphotransacetylase from the methanogenic archaeon Methanosarcina thermophila in complex with the substrate CoA revealed one CoA (CoA1) bound in the proposed active site cleft and an additional CoA (CoA2) bound at the periphery of the cleft. The results of isothermal titration calorimetry experiments are described, and they support the hypothesis that there are distinct high-affinity (equilibrium dissociation constant [KD], 20 microM) and low-affinity (KD, 2 mM) CoA binding sites. The crystal structures indicated that binding of CoA1 is mediated by a series of hydrogen bonds and extensive van der Waals interactions with the enzyme and that there are fewer of these interactions between CoA2 and the enzyme. Different conformations of the protein observed in the crystal structures suggest that domain movements which alter the geometry of the active site cleft may contribute to catalysis. Kinetic and calorimetric analyses of site-specific replacement variants indicated that there are catalytic roles for Ser309 and Arg310, which are proximal to the reactive sulfhydryl of CoA1. The reaction is hypothesized to proceed through base-catalyzed abstraction of the thiol proton of CoA by the adjacent and invariant residue Asp316, followed by nucleophilic attack of the thiolate anion of CoA on the carbonyl carbon of acetyl phosphate. We propose that Arg310 binds acetyl phosphate and orients it for optimal nucleophilic attack. The hypothesized mechanism proceeds through a negatively charged transition state stabilized by hydrogen bond donation from Ser309.
Organizational Affiliation:
Department of Biochemistry, Center for Structural Biology, SUNY Stony Brook, Stony Brook, NY 11794-5115, USA.