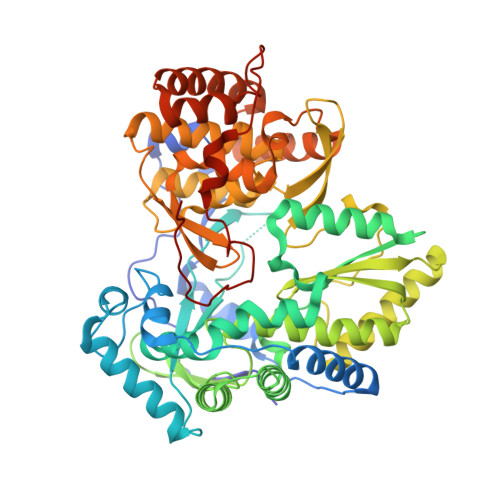
Structure of Hepatitis C Virus Ns5B Polymerase in a New Crystal Form: Insights Into Oligomerisation and Allosteric Nucleotide Binding Site
Biswal, B.K., Cherney, M.M., Chan, L., Yannopoulos, C.G., Bilimoria, D., Bedard, J., James, M.N.G.To be published.