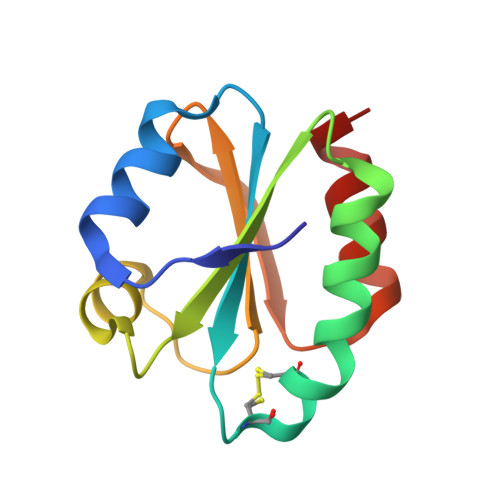
Conservation of protein structure over four billion years.
Ingles-Prieto, A., Ibarra-Molero, B., Delgado-Delgado, A., Perez-Jimenez, R., Fernandez, J.M., Gaucher, E.A., Sanchez-Ruiz, J.M., Gavira, J.A.(2013) Structure 21: 1690-1697
- PubMed: 23932589
- DOI: https://doi.org/10.1016/j.str.2013.06.020
- Primary Citation of Related Structures:
2YJ7, 2YN1, 2YNX, 2YOI, 2YPM, 3ZIV, 4BA7 - PubMed Abstract:
Little is known about the evolution of protein structures and the degree of protein structure conservation over planetary time scales. Here, we report the X-ray crystal structures of seven laboratory resurrections of Precambrian thioredoxins dating up to approximately four billion years ago. Despite considerable sequence differences compared with extant enzymes, the ancestral proteins display the canonical thioredoxin fold, whereas only small structural changes have occurred over four billion years. This remarkable degree of structure conservation since a time near the last common ancestor of life supports a punctuated-equilibrium model of structure evolution in which the generation of new folds occurs over comparatively short periods and is followed by long periods of structural stasis.
Organizational Affiliation:
Departamento de Química Física, Facultad de Ciencias, Universidad de Granada, Granada 18071, Spain.