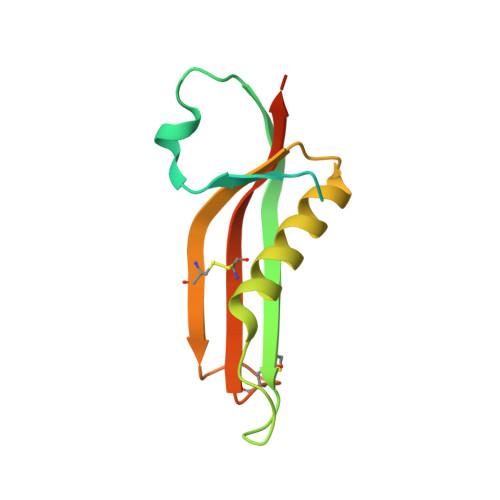
Crystal Structure of the Outer Membrane Protein Rcsf, a New Substrate for the Periplasmic Protein- Disulfide Isomerase Dsbc.
Leverrier, P., Declercq, J.P., Denoncin, K., Vertommen, D., Hiniker, A., Cho, S.H., Collet, J.F.(2011) J Biol Chem 286: 16734
- PubMed: 21454485
- DOI: https://doi.org/10.1074/jbc.M111.224865
- Primary Citation of Related Structures:
2Y1B - PubMed Abstract:
The bacterial Rcs phosphorelay is a stress-induced defense mechanism that controls the expression of numerous genes, including those for capsular polysaccharides, motility, and virulence factors. It is a complex multicomponent system that includes the histidine kinase (RcsC) and the response regulator (RcsB) and also auxiliary proteins such as RcsF. RcsF is an outer membrane lipoprotein that transmits signals from the cell surface to RcsC. The physiological signals that activate RcsF and how RcsF interacts with RcsC remain unknown. Here, we report the three-dimensional structure of RcsF. The fold of the protein is characterized by the presence of a central 4-stranded β sheet, which is conserved in several other proteins, including the copper-binding domain of the amyloid precursor protein. RcsF, which contains four conserved cysteine residues, presents two nonconsecutive disulfides between Cys(74) and Cys(118) and between Cys(109) and Cys(124), respectively. These two disulfides are not functionally equivalent; the Cys(109)-Cys(124) disulfide is particularly important for the assembly of an active RcsF. Moreover, we show that formation of the nonconsecutive disulfides of RcsF depends on the periplasmic disulfide isomerase DsbC. We trapped RcsF in a mixed disulfide complex with DsbC, and we show that deletion of dsbC prevents the activation of the Rcs phosphorelay by signals that function through RcsF. The three-dimensional structure of RcsF provides the structural basis to understand how this protein triggers the Rcs signaling cascade.
Organizational Affiliation:
Welbio (Walloon Excellence in Life Sciences and Biotechnology), Université Catholique de Louvain, B-1200 Brussels, Belgium.