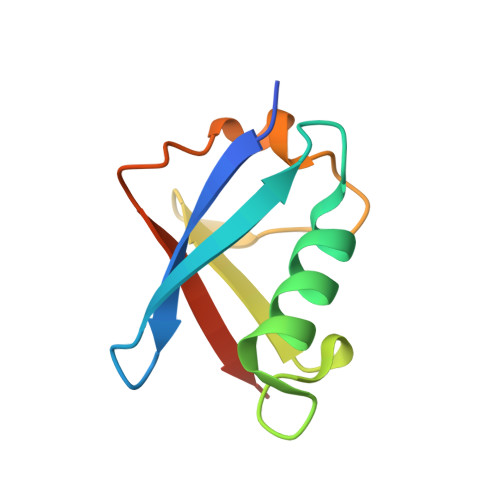
The Crystal Structure of the Ubiquitin-Like (Ubl) Domain of Human Homologue a of Rad23 (Hhr23A) Protein
Chen, Y.W., Tajima, T., Agrawal, S.(2011) Protein Eng Des Sel 24: 131
- PubMed: 21047872
- DOI: https://doi.org/10.1093/protein/gzq084
- Primary Citation of Related Structures:
2WYQ - PubMed Abstract:
The human homologue of the yeast Rad23 protein, hHR23A, plays dual roles in DNA repair as well as in translocating polyubiquitinated proteins to the proteasome. We determined the three-dimensional structure of its ubiquitin-like (UbL) domain by X-ray crystallography. It has the same overall structure and fold characteristics as ubiquitin and other members of the UbL domain family, with overall root mean square deviations in Cα positions in the range of 1.0-1.3 Å. There are local differences in the α1-β3 loop where hHR23A UbL domain has three more residues constituting a bigger loop. Analysis of the crystal packing revealed a possible dimeric arrangement mediated by the three residues (Leu10, Ile49 and Met75) that are known to be critical for molecular interactions. In contrast to the overall well-defined structure, these three residues are either disordered or have multiple conformations, suggesting that conformation variability is an important property of the binding surface. The electrostatic potentials at the binding surface are conserved among the family, with the hHR23B domain being the most similar to this structure. The intra-molecular complexes formed by the UbL domain of hHR23A with its UbA1 or UbA2 domains was studied by comparative homology modelling, which suggests these two interactions are structurally similar and are mutually exclusive.
Organizational Affiliation:
King's College London, Randall Division of Cell and Molecular Biophysics, New Hunt's House, Guy's Campus, London SE11UL, UK. yu-wai.chen@kcl.ac.uk