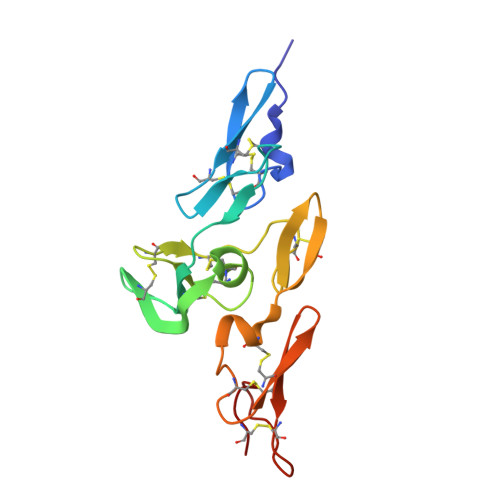
Structure and Interdomain Interactions of a Hybrid Domain: A Disulphide-Rich Module of the Fibrillin/Ltbp Superfamily of Matrix Proteins.
Jensen, S.A., Iqbal, S., Lowe, E.D., Redfield, C., Handford, P.A.(2009) Structure 17: 759
- PubMed: 19446531
- DOI: https://doi.org/10.1016/j.str.2009.03.014
- Primary Citation of Related Structures:
2W86 - PubMed Abstract:
The fibrillins and latent transforming growth factor-beta binding proteins (LTBPs) form a superfamily of structurally-related proteins consisting of calcium-binding epidermal growth factor-like (cbEGF) domains interspersed with 8-cysteine-containing transforming growth factor beta-binding protein-like (TB) and hybrid (hyb) domains. Fibrillins are the major components of the extracellular 10-12 nm diameter microfibrils, which mediate a variety of cell-matrix interactions. Here we present the crystal structure of a fibrillin-1 cbEGF9-hyb2-cbEGF10 fragment, solved to 1.8 A resolution. The hybrid domain fold is similar, but not identical, to the TB domain fold seen in previous fibrillin-1 and LTBP-1 fragments. Pairwise interactions with neighboring cbEGF domains demonstrate extensive interfaces, with the hyb2-cbEGF10 interface dependent on Ca(2+) binding. These observations provide accurate constraints for models of fibrillin organization within the 10-12 nm microfibrils and provide further molecular insights into how Ca(2+) binding influences the intermolecular interactions and biomechanical properties of fibrillin-1.
Organizational Affiliation:
Department of Biochemistry, University of Oxford, Oxford, UK.