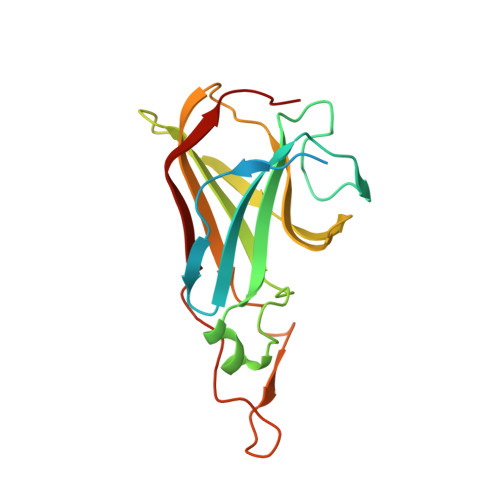
Crystal structure of the Tp34 (TP0971) lipoprotein of treponema pallidum: implications of its metal-bound state and affinity for human lactoferrin.
Deka, R.K., Brautigam, C.A., Tomson, F.L., Lumpkins, S.B., Tomchick, D.R., Machius, M., Norgard, M.V.(2007) J Biol Chem 282: 5944-5958
- PubMed: 17192261
- DOI: https://doi.org/10.1074/jbc.M610215200
- Primary Citation of Related Structures:
2O6C, 2O6D, 2O6E, 2O6F - PubMed Abstract:
The Tp34 (TP0971) membrane lipoprotein of Treponema pallidum, an obligate human pathogen and the agent of syphilis, was previously reported to have lactoferrin binding properties. Given the non-cultivatable nature of T. pallidum, a structure-to-function approach was pursued to clarify further potential relationships between the Tp34 structural and biochemical properties and its propensity to bind human lactoferrin. The crystal structure of a nonacylated, recombinant form of Tp34 (rTp34), solved to a resolution of 1.9A(,) revealed two metaloccupied binding sites within a dimer; the identity of the ion most likely was zinc. Residues from both of the monomers contributed to the interfacial metal-binding sites; a novel feature was that the delta-sulfur of methionine coordinated the zinc ion. Analytical ultracentrifugation showed that, in solution, rTp34 formed a metal-stabilized dimer and that rTp34 bound human lactoferrin with a stoichiometry of 2:1. Isothermal titration calorimetry further revealed that rTp34 bound human lactoferrin at high (submicromolar) affinity. Finally, membrane topology studies revealed that native Tp34 is not located on the outer surface (outer membrane) of T. pallidum but, rather, is periplasmic. How propensity of Tp34 to bind zinc and the iron-sequestering lactoferrin may relate overall to the biology of T. pallidum infection in humans is discussed.
Organizational Affiliation:
Departments of Microbiology and Biochemistry, University of Texas Southwestern Medical Center, Dallas, Texas 75390, USA.