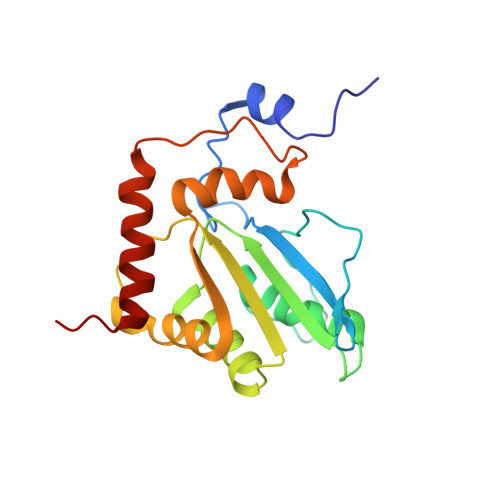
Structure, interaction and real-time monitoring of the enzymatic reaction of wild-type APOBEC3G
Furukawa, A., Nagata, T., Matsugami, A., Habu, Y., Sugiyama, R., Hayashi, F., Kobayashi, N., Yokoyama, S., Takaku, H., Katahira, M.(2009) EMBO J 28: 440-451
- PubMed: 19153609
- DOI: https://doi.org/10.1038/emboj.2008.290
- Primary Citation of Related Structures:
2KBO - PubMed Abstract:
Human APOBEC3G exhibits anti-human immunodeficiency virus-1 (HIV-1) activity by deaminating cytidines of the minus strand of HIV-1. Here, we report a solution structure of the C-terminal deaminase domain of wild-type APOBEC3G. The interaction with DNA was examined. Many differences in the interaction were found between the wild type and recently studied mutant APOBEC3Gs. The position of the substrate cytidine, together with that of a DNA chain, in the complex, was deduced. Interestingly, the deamination reaction of APOBEC3G was successfully monitored using NMR signals in real time. Real-time monitoring has revealed that the third cytidine of the d(CCCA) segment is deaminated at an early stage and that then the second one is deaminated at a late stage, the first one not being deaminated at all. This indicates that the deamination is carried out in a strict 3' --> 5' order. Virus infectivity factor (Vif) of HIV-1 counteracts the anti-HIV-1 activity of APOBEC3G. The structure of the N-terminal domain of APOBEC3G, with which Vif interacts, was constructed with homology modelling. The structure implies the mechanism of species-specific sensitivity of APOBEC3G to Vif action.
Organizational Affiliation:
Supramolecular Biology, International Graduate School of Arts and Sciences, Yokohama City University, Yokohama, Japan.