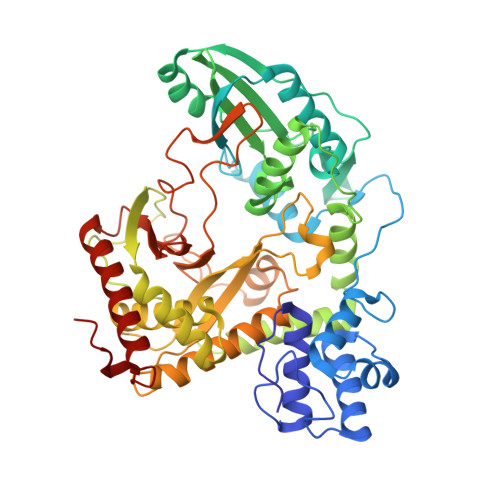
Structural and Functional Insights Into a Peptide Bond-Forming Bidomain from a Nonribosomal Peptide Synthetase.
Samel, S.A., Schoenafinger, G., Knappe, T.A., Marahiel, M.A., Essen, L.-O.(2007) Structure 15: 781
- PubMed: 17637339
- DOI: https://doi.org/10.1016/j.str.2007.05.008
- Primary Citation of Related Structures:
2JGP - PubMed Abstract:
The crystal structure of the bidomain PCP-C from modules 5 and 6 of the nonribosomal tyrocidine synthetase TycC was determined at 1.8 A resolution. The bidomain structure reveals a V-shaped condensation domain, the canyon-like active site groove of which is associated with the preceding peptidyl carrier protein (PCP) domain at its donor side. The relative arrangement of the PCP and the peptide bond-forming condensation (C) domain places the active sites approximately 50 A apart. Accordingly, this PCP-C structure represents a conformational state prior to peptide transfer from the donor-PCP to the acceptor-PCP domain, implying the existence of additional states of PCP-C domain interaction during catalysis. Additionally, PCP-C exerts a mode of cyclization activity that mimics peptide bond formation catalyzed by C domains. Based on mutational data and pK value analysis of active site residues, it is suggested that nonribosomal peptide bond formation depends on electrostatic interactions rather than on general acid/base catalysis.
Organizational Affiliation:
Department of Chemistry/Biochemistry, Philipps-Universität, Hans-Meerwein-Strasse, D-35032 Marburg, Germany.