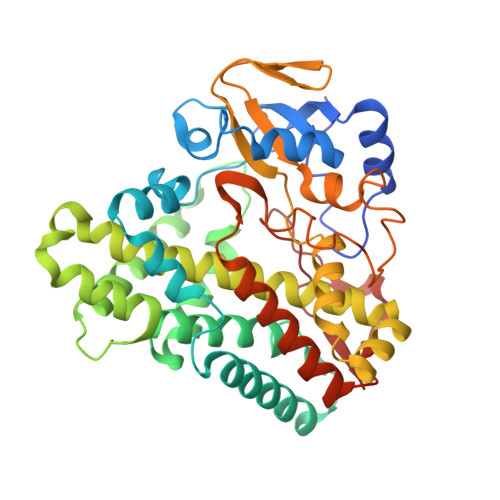
Crystal structure of the Mycobacterium tuberculosis P450 CYP121-fluconazole complex reveals new azole drug-P450 binding mode.
Seward, H.E., Roujeinikova, A., McLean, K.J., Munro, A.W., Leys, D.(2006) J Biol Chem 281: 39437-39443
- PubMed: 17028183
- DOI: https://doi.org/10.1074/jbc.M607665200
- Primary Citation of Related Structures:
2IJ5, 2IJ7 - PubMed Abstract:
Azole and triazole drugs are cytochrome P450 inhibitors widely used as fungal antibiotics and possessing potent antimycobacterial activity. We present here the crystal structure of Mycobacterium tuberculosis cytochrome P450 CYP121 in complex with the triazole drug fluconazole, revealing a new azole heme ligation mode. In contrast to other structurally characterized cytochrome P450 azole complexes, where the azole nitrogen directly coordinates the heme iron, in CYP121 fluconazole does not displace the aqua sixth heme ligand but occupies a position that allows formation of a direct hydrogen bond to the aqua sixth heme ligand. Direct ligation of fluconazole to the heme iron is observed in a minority of CYP121 molecules, albeit with severe deviations from ideal geometry due to close contacts with active site residues. Analysis of both ligand-on and -off structures reveals the relative position of active site residues derived from the I-helix is a key determinant in the relative ratio of on and off states. Regardless, both ligand-bound states lead to P450 inactivation by active site occlusion. This previously unrecognized means of P450 inactivation is consistent with spectroscopic analyses in both solution and in the crystalline form and raises important questions relating to interaction of azoles with both pathogen and human P450s.
Organizational Affiliation:
Department of Biochemistry, University of Leicester, Henry Wellcome Building, Lancaster Road, Leicester LE6 0HQ, United Kingdom.