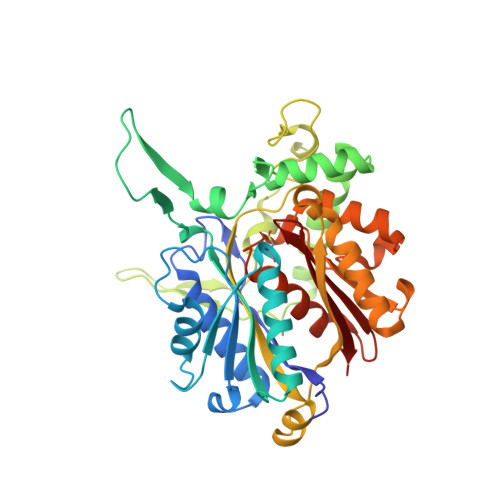
Crystallographic and Kinetic Studies of Human Mitochondrial Acetoacetyl-CoA Thiolase: The Importance of Potassium and Chloride Ions for Its Structure and Function
Haapalainen, A.M., Merilainen, G., Pirila, P.L., Kondo, N., Fukao, T., Wierenga, R.K.(2007) Biochemistry 46: 4305-4321
- PubMed: 17371050
- DOI: https://doi.org/10.1021/bi6026192
- Primary Citation of Related Structures:
2IB7, 2IB8, 2IB9, 2IBU, 2IBW, 2IBY - PubMed Abstract:
Thiolases are CoA-dependent enzymes which catalyze the formation of a carbon-carbon bond in a Claisen condensation step and its reverse reaction via a thiolytic degradation mechanism. Mitochondrial acetoacetyl-coenzyme A (CoA) thiolase (T2) is important in the pathways for the synthesis and degradation of ketone bodies as well as for the degradation of 2-methylacetoacetyl-CoA. Human T2 deficiency has been identified in more than 60 patients. A unique property of T2 is its activation by potassium ions. High-resolution human T2 crystal structures are reported for the apo form and the CoA complex, with and without a bound potassium ion. The potassium ion is bound near the CoA binding site and the catalytic site. Binding of the potassium ion at this low-affinity binding site causes the rigidification of a CoA binding loop and an active site loop. Unexpectedly, a high-affinity binding site for a chloride ion has also been identified. The chloride ion is copurified, and its binding site is at the dimer interface, near two catalytic loops. A unique property of T2 is its ability to use 2-methyl-branched acetoacetyl-CoA as a substrate, whereas the other structurally characterized thiolases cannot utilize the 2-methylated compounds. The kinetic measurements show that T2 can degrade acetoacetyl-CoA and 2-methylacetoacetyl-CoA with similar catalytic efficiencies. For both substrates, the turnover numbers increase approximately 3-fold when the potassium ion concentration is increased from 0 to 40 mM KCl. The structural analysis of the active site of T2 indicates that the Phe325-Pro326 dipeptide near the catalytic cavity is responsible for the exclusive 2-methyl-branched substrate specificity.
Organizational Affiliation:
Biocenter Oulu and Department of Biochemistry, University of Oulu, P.O. Box 3000, FIN-90014 Oulu, Finland.