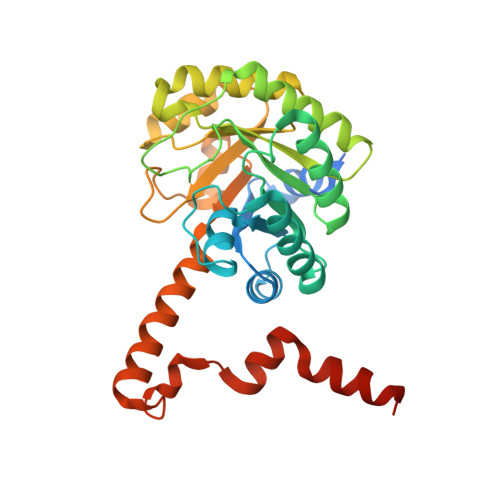
Crystal Structure of the Petal Death Protein from Carnation Flower.
Teplyakov, A., Liu, S., Lu, Z., Howard, A., Dunaway-Mariano, D., Herzberg, O.(2005) Biochemistry 44: 16377-16384
- PubMed: 16342930
- DOI: https://doi.org/10.1021/bi051779y
- Primary Citation of Related Structures:
1ZLP - PubMed Abstract:
Expression of the PSR132 protein from Dianthus caryophyllus (carnation, clover pink) is induced in response to ethylene production associated with petal senescence, and thus the protein is named petal death protein (PDP). Recent work has established that despite the annotation of PDP in sequence databases as carboxyphosphoenolpyruvate mutase, the enzyme is actually a C-C bond cleaving lyase exhibiting a broad substrate profile. The crystal structure of PDP has been determined at 2.7 A resolution, revealing a dimer-of-dimers oligomeric association. Consistent with sequence homology, the overall alpha/beta barrel fold of PDP is the same as that of other isocitrate lyase/PEP mutase superfamily members, including a swapped eighth helix within a dimer. Moreover, Mg(2+) binds in the active site of PDP with a coordination pattern similar to that seen in other superfamily members. A compound, covalently bound to the catalytic residue, Cys144, was interpreted as a thiohemiacetal adduct resulting from the reaction of glutaraldehyde used to cross-link the crystals. The Cys144-carrying flexible loop that gates access to the active site is in the closed conformation. Models of bound substrates and comparison with the closed conformation of isocitrate lyase and 2-methylisocitrate lyase revealed the structural basis for the broad substrate profile of PDP.
Organizational Affiliation:
Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute, Rockville, Maryland 20850, USA.