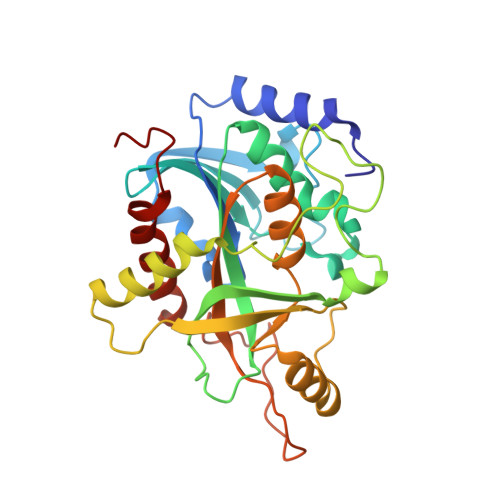
Kinetics and crystal structure of human purine nucleoside phosphorylase in complex with 7-methyl-6-thio-guanosine
Silva, R.G., Pereira, J.H., Canduri, F., de Azevedo Jr., W.F., Basso, L.A., Santos, D.S.(2005) Arch Biochem Biophys 442: 49-58
- PubMed: 16154528
- DOI: https://doi.org/10.1016/j.abb.2005.07.021
- Primary Citation of Related Structures:
1YRY - PubMed Abstract:
Purine nucleoside phosphorylase (PNP) catalyzes the reversible phosphorolysis of nucleosides and deoxynucleosides, generating ribose 1-phosphate and the purine base, which is an important step of purine catabolism pathway. The lack of such an activity in humans, owing to a genetic disorder, causes T-cell impairment, and drugs that inhibit this enzyme may have the potential of being utilized as modulators of the immunological system to treat leukemia, autoimmune diseases, and rejection in organ transplantation. Here, we describe kinetics and crystal structure of human PNP in complex with 7-methyl-6-thio-guanosine, a synthetic substrate, which is largely used in activity assays. Analysis of the structure identifies different protein conformational changes upon ligand binding, and comparison of kinetic and structural data permits an understanding of the effects of atomic substitution on key positions of the synthetic substrate and their consequences to enzyme binding and catalysis. Such knowledge may be helpful in designing new PNP inhibitors.
Organizational Affiliation:
Centro de Pesquisas em Biologia Molecular e Funcional, Instituto de Pesquisas Biomédicas, PUCRS, Porto Alegre, RS, Brazil.