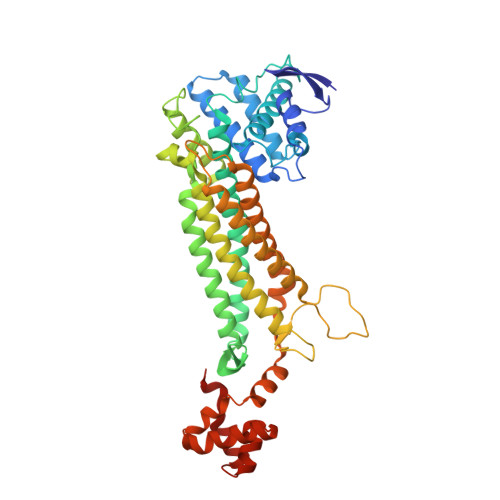
Structure of free fumarase C from Escherichia coli.
Weaver, T.(2005) Acta Crystallogr D Biol Crystallogr 61: 1395-1401
- PubMed: 16204892
- DOI: https://doi.org/10.1107/S0907444905024194
- Primary Citation of Related Structures:
1YFE - PubMed Abstract:
Previous crystal structures of fumarase C from Escherichia coli have noted two occupied dicarboxylate-binding sites termed the active site and the B site. Here, the first known fumarase C structure is reported in which both sites are unoccupied by bound ligand. This so-called ;free' crystal form shows conservation of the active-site water in a similar orientation to that reported in other fumarase C crystal structures. More importantly, a shift of His129 has been observed at the B site. This new crystallographic information suggests the use of water as a permanent member of the active site and the use of an imidazole-imidazolium conversion to control access at the allosteric B site.
Organizational Affiliation:
Department of Chemistry, University of Wisconsin-La Crosse, 1725 State Street, 4020 Cowley Hall, La Crosse, Wisconsin 54601, USA. weaver.todd@uwlax.edu