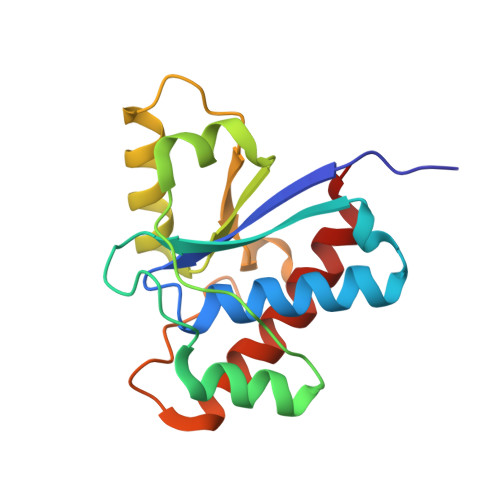
Crystal structure of the human B-form low molecular weight phosphotyrosyl phosphatase at 1.6-A resolution.
Zabell, A.P., Schroff, A.D., Bain, B.E., Van Etten, R.L., Wiest, O., Stauffacher, C.V.(2006) J Biol Chem 281: 6520-6527
- PubMed: 16253994
- DOI: https://doi.org/10.1074/jbc.M506285200
- Primary Citation of Related Structures:
1XWW - PubMed Abstract:
The crystal structure of HPTP-B, a human isoenzyme of the low molecular weight phosphotyrosyl phosphatase (LMW PTPase) is reported here at a resolution of 1.6 A. This high resolution structure of the second human LMW PTPase isoenzyme provides the opportunity to examine the structural basis of different substrate and inhibitor/activator responses. The crystal packing of HPTP-B positions a normally surface-exposed arginine in a position equivalent to the tyrosyl substrate. A comparison of all deposited crystallographic coordinates of these PTPases reveals three atomic positions within the active site cavity occupied by hydrogen bond donor or acceptor atoms on bound molecules, suggesting useful design elements for synthetic inhibitors. A selection of inhibitor and activator molecules as well as small molecule and peptide substrates were tested against each human isoenzyme. These results along with the crystal packing seen in HPTP-B suggest relevant sequence elements in the currently unknown target sequence.
Organizational Affiliation:
Department of Biological Sciences and the Purdue Cancer Center, Purdue University, West Lafayette, Indiana 47907-1392, USA.