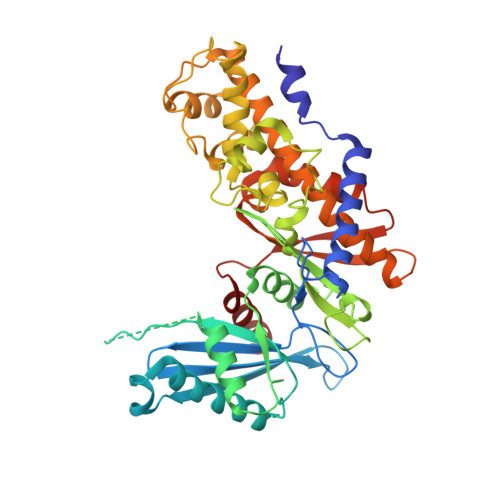
Structural basis for allosteric regulation of the monomeric allosteric enzyme human glucokinase
Kamata, K., Mitsuya, M., Nishimura, T., Eiki, J., Nagata, Y.(2004) Structure 12: 429-438
- PubMed: 15016359
- DOI: https://doi.org/10.1016/j.str.2004.02.005
- Primary Citation of Related Structures:
1V4S, 1V4T - PubMed Abstract:
Glucokinase is a monomeric enzyme that displays a low affinity for glucose and a sigmoidal saturation curve for its substrate, two properties that are important for its playing the role of a glucose sensor in pancreas and liver. The molecular basis for these two properties is not well understood. Herein we report the crystal structures of glucokinase in its active and inactive forms, which demonstrate that global conformational change, including domain reorganization, is induced by glucose binding. This suggests that the positive cooperativity of monomeric glucokinase obeys the "mnemonical mechanism" rather than the well-known concerted model. These structures also revealed an allosteric site through which small molecules may modulate the kinetic properties of the enzyme. This finding provided the mechanistic basis for activation of glucokinase as a potential therapeutic approach for treating type 2 diabetes mellitus.
Organizational Affiliation:
Banyu Tsukuba Research Institute in collaboration with Merck Research Laboratories, Okubo 3, Tsukuba, Ibaraki 300-2611, Japan. kenji_kamata@merck.com