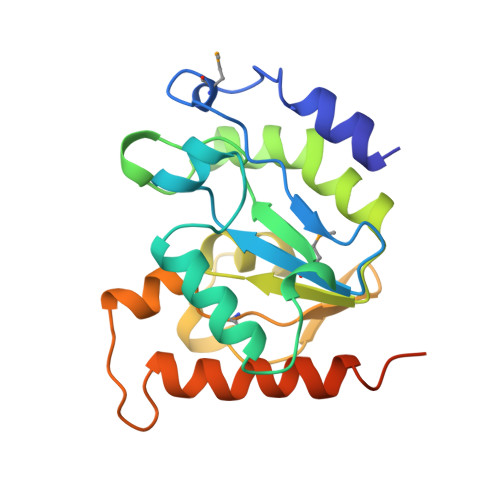
Crystal Structure of a Family 4 Uracil-DNA Glycosylase from Thermus thermophilus HB8
Hoseki, J., Okamoto, A., Masui, R., Shibata, T., Inoue, Y., Yokoyama, S., Kuramitsu, S.(2003) J Mol Biol 333: 515-526
- PubMed: 14556741
- DOI: https://doi.org/10.1016/j.jmb.2003.08.030
- Primary Citation of Related Structures:
1UI0, 1UI1 - PubMed Abstract:
Uracil-DNA glycosylase (UDG; EC 3.2.2.-) removes uracil from DNA to initiate DNA base excision repair. Since hydrolytic deamination of cytosine to uracil is one of the most frequent DNA-damaging events in all cells, UDG is an essential enzyme for maintaining the integrity of genomic information. For the first time, we report the crystal structure of a family 4 UDG from Thermus thermophilus HB8 (TthUDG) complexed with uracil, solved at 1.5 angstroms resolution. As opposed to UDG enzymes in its other families, TthUDG possesses a [4Fe-4S] cluster. This iron-sulfur cluster, which is distant from the active site, interacts with loop structures and has been suggested to be unessential to the activity but necessary for stabilizing the loop structures. In addition to the iron-sulfur cluster, salt-bridges and ion pairs on the molecular surface and the presence of proline on loops and turns is thought to contribute to the enzyme's thermostability. Despite very low levels of sequence identity with Escherichia coli and human UDGs (family 1) and E.coli G:T/U mismatch-specific DNA glycosylase (MUG) (family 2), the topology and order of secondary structures of TthUDG are similar to those of these distant relatives. Furthermore, the coordinates of the core structure formed by beta-strands are almost the same. Positive charge is distributed over the active-site groove, where TthUDG would bind DNA strands, as do UDG enzymes in other families. TthUDG recognizes uracil specifically in the same manner as does human UDG (family 1), rather than guanine in the complementary strand DNA, as does E.coli MUG (family 2). These results suggest that the mechanism by which family 4 UDGs remove uracils from DNA is similar to that of family 1 enzymes.
Organizational Affiliation:
RIKEN Harima Institute at SPring-8, 1-1-1 Kouto, Mikazuki, Sayo-gun, Hyogo 679-5148, Japan.