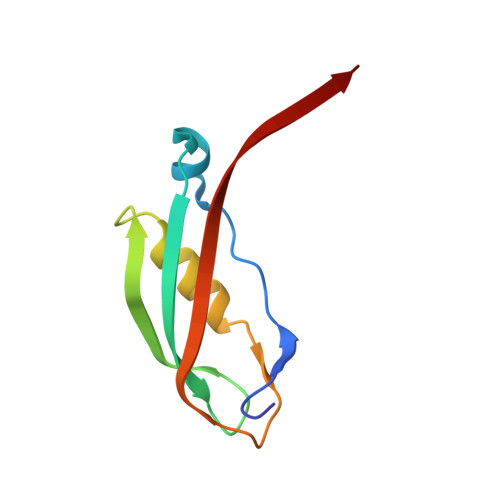
Crystal structure of a 92-residue c-terminal fragment of TonB from Escherichia coli reveals significant conformational changes compared to structures of smaller TonB fragments
Koedding, J., Killig, F., Polzer, P., Howard, S.P., Diederichs, K., Welte, W.(2005) J Biol Chem 280: 3022-3028
- PubMed: 15522863
- DOI: https://doi.org/10.1074/jbc.M411155200
- Primary Citation of Related Structures:
1U07 - PubMed Abstract:
Uptake of siderophores and vitamin B(12) through the outer membrane of Escherichia coli is effected by an active transport system consisting of several outer membrane receptors and a protein complex of the inner membrane. The link between these is TonB, a protein associated with the cytoplasmic membrane, which forms a large periplasmic domain capable of interacting with several outer membrane receptors, e.g. FhuA, FecA, and FepA for siderophores and BtuB for vitamin B(12.) The active transport across the outer membrane is driven by the chemiosmotic gradient of the inner membrane and is mediated by the TonB protein. The receptor-binding domain of TonB appears to be formed by a highly conserved C-terminal amino acid sequence of approximately 100 residues. Crystal structures of two C-terminal TonB fragments composed of 85 (TonB-85) and 77 (TonB-77) amino acid residues, respectively, have been previously determined (Chang, C., Mooser, A., Pluckthun, A., and Wlodawer, A. (2001) J. Biol. Chem. 276, 27535-27540 and Koedding, J., Howard, S. P., Kaufmann, L., Polzer, P., Lustig, A., and Welte, W. (2004) J. Biol. Chem. 279, 9978-9986). In both cases the TonB fragments form dimers in solution and crystallize as dimers consisting of monomers tightly engaged with one another by the exchange of a beta-hairpin and a C-terminal beta-strand. Here we present the crystal structure of a 92-residue fragment of TonB (TonB-92), which is monomeric in solution. The structure, determined at 1.13-A resolution, shows a dimer with considerably reduced intermolecular interaction compared with the other known TonB structures, in particular lacking the beta-hairpin exchange.
Organizational Affiliation:
Department of Biology, University of Konstanz, Universitätsstrasse 10, 78457 Konstanz, Germany.