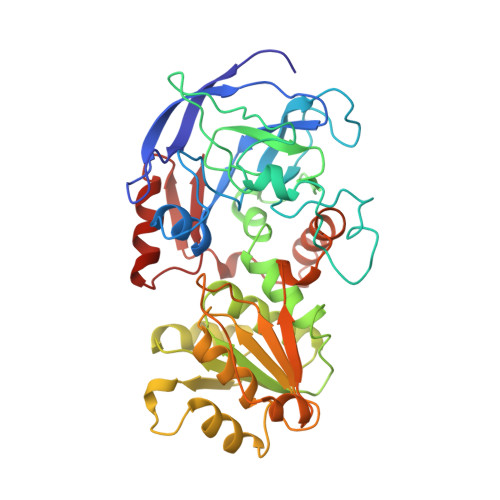
Structure of human chi chi alcohol dehydrogenase: a glutathione-dependent formaldehyde dehydrogenase.
Yang, Z.N., Bosron, W.F., Hurley, T.D.(1997) J Mol Biol 265: 330-343
- PubMed: 9018047
- DOI: https://doi.org/10.1006/jmbi.1996.0731
- Primary Citation of Related Structures:
1TEH - PubMed Abstract:
The crystal structure of the human class III chi chi alcohol dehydrogenase (ADH) in a binary complex with NAD+(gamma) was solved to 2.7 A resolution by molecular replacement with human class I beta1 beta1 ADH. chi chi ADH catalyzes the oxidation of long-chain alcohols such as omega-hydroxy fatty acids as well as S-hydroxymethyl-glutathione, a spontaneous adduct between formaldehyde and glutathione. There are two subunits per asymmetric unit in the chi chi ADH structure. Both subunits display a semi-open conformation of the catalytic domain. This conformation is half-way between the open and closed conformations described for the horse EE ADH enzyme. The semi-open conformation and key changes in elements of secondary structure provide a structural basis for the ability of chi chi ADH to bind S-hydroxymethyl-glutathione and 10-hydroxydecanoate. Direct coordination of the catalytic zinc ion by Glu68 creates a novel environment for the catalytic zinc ion in chi chi ADH. This new configuration of the catalytic zinc is similar to an intermediate for horse EE ADH proposed through theoretical computations and is consistent with the spectroscopic data of the Co(II)-substituted chi chi enzyme. The position for residue His47 in the chi chi ADH structure suggests His47 may function both as a catalytic base for proton transfer and in the binding of the adenosine phosphate of NAD(H). Modeling of substrate binding to this enzyme structure is consistent with prior mutagenesis data which showed that both Asp57 and Arg115 contribute to glutathione binding and that Arg115 contributes to the binding of omega-hydroxy fatty acids and identifies additional residues which may contribute to substrate binding.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, Indianapolis 46202, USA.