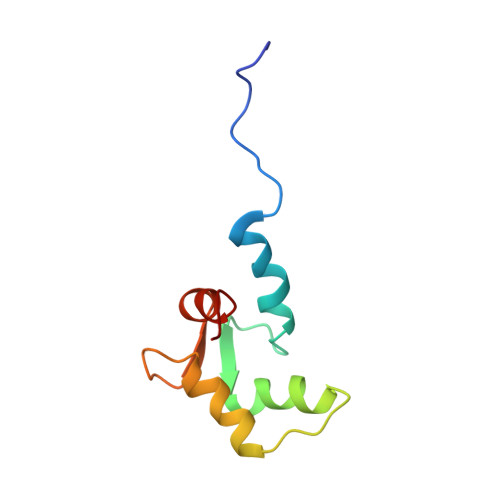
Structure of the Mg2+-loaded C-lobe of cardiac troponin C bound to the N-domain of cardiac troponin I: comparison with the Ca2+-loaded structure.
Finley, N.L., Howarth, J.W., Rosevear, P.R.(2004) Biochemistry 43: 11371-11379
- PubMed: 15350124
- DOI: https://doi.org/10.1021/bi049672i
- Primary Citation of Related Structures:
1SBJ, 1SCV - PubMed Abstract:
Cardiac troponin C (cTnC) is the Ca(2+)-binding component of the troponin complex and, as such, is the Ca(2+)-dependent switch in muscle contraction. This protein consists of two globular lobes, each containing a pair of EF-hand metal-binding sites, connected by a linker. In the N lobe, Ca(2+)-binding site I is inactive and Ca(2+)-binding site II is primarily responsible for initiation of muscle contraction. The C lobe contains Ca(2+)/Mg(2+)-binding sites III and IV, which bind Mg(2+) with lower affinity and play a structural as well as a secondary role in modulating the Ca(2+) signal. To understand the structural consequences of Ca(2+)/Mg(2+) exchange in the C lobe, we have determined the NMR solution structure of the Mg(2+)-loaded C lobe, cTnC(81-161), in a complex with the N domain of cardiac troponin I, cTnI(33-80), and compared it with a refined Ca(2+)-loaded structure. The overall tertiary structure of the Mg(2+)-loaded C lobe is very similar to that of the refined Ca(2+)-loaded structure as evidenced by the root-mean-square deviation of 0.94 A for all backbone atoms. While metal-dependent conformational changes are minimal, substitution of Mg(2+) for Ca(2+) is characterized by condensation of the C-terminal portion of the metal-binding loops with monodentate Mg(2+) ligation by the conserved Glu at position 12 and partial closure of the cTnI hydrophobic binding cleft around site IV. Thus, conformational plasticity in the Ca(2+)/Mg(2+)-dependent binding loops may represent a mechanism to modulate C-lobe cTnC interactions with the N domain of cTnI.
Organizational Affiliation:
Department of Molecular Genetics, Biochemistry, and Microbiology, University of Cincinnati, College of Medicine, 231 Albert Sabin Way, Medical Sciences Building, Cincinnati Ohio 45267-0524, USA.