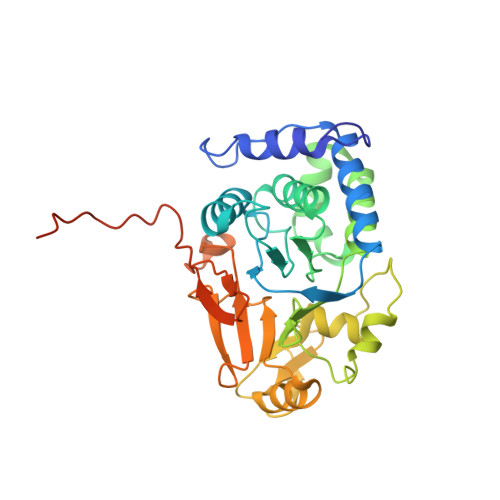
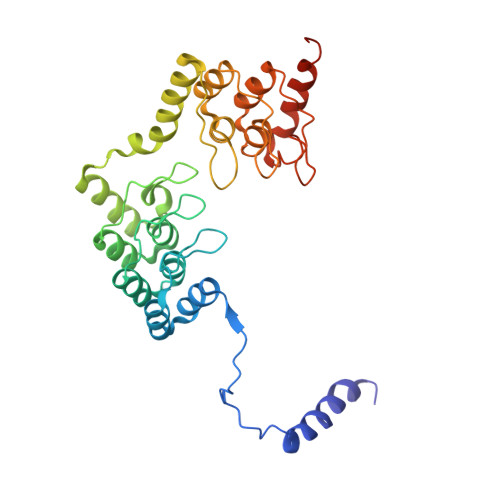
Structural basis of protein phosphatase 1 regulation
Terrak, M., Kerff, F., Langsetmo, K., Tao, T., Dominguez, R.(2004) Nature 429: 780-784
- PubMed: 15164081
- DOI: https://doi.org/10.1038/nature02582
- Primary Citation of Related Structures:
1S70 - PubMed Abstract:
The coordinated and reciprocal action of serine/threonine (Ser/Thr) protein kinases and phosphatases produces transient phosphorylation, a fundamental regulatory mechanism for many biological processes. The human genome encodes a far greater number of Ser/Thr protein kinases than of phosphatases. Protein phosphatase 1 (PP1), in particular, is ubiquitously distributed and regulates a broad range of cellular functions, including glycogen metabolism, cell-cycle progression and muscle relaxation. PP1 has evolved effective catalytic machinery but lacks substrate specificity. Substrate specificity is conferred upon PP1 through interactions with a large number of regulatory subunits. The regulatory subunits are generally unrelated, but most possess the RVxF motif, a canonical PP1-binding sequence. Here we reveal the crystal structure at 2.7 A resolution of the complex between PP1 and a 34-kDa N-terminal domain of the myosin phosphatase targeting subunit MYPT1. MYPT1 is the protein that regulates PP1 function in smooth muscle relaxation. Structural elements amino- and carboxy-terminal to the RVxF motif of MYPT1 are positioned in a way that leads to a pronounced reshaping of the catalytic cleft of PP1, contributing to the increased myosin specificity of this complex. The structure has general implications for the control of PP1 activity by other regulatory subunits.
Organizational Affiliation:
Boston Biomedical Research Institute, 64 Grove Street, Watertown, Massachusetts 02472, USA.